A Comprehensive Workflow for Biosimilar Comparability Using Peptide Mapping Analysis with Trastuzumab
Abstract
This application note describes a complete LC-MS workflow for peptide mapping analysis of monoclonal antibodies (mAbs) to streamline biosimilarity assessment, demonstrated through the comparison of an innovator biologic, Trastuzumab, with a biosimilar.
The workflow provides the following:
- A detailed protocol for filter assisted sample preparation (FASP), for peptide mapping with minimal protocol-induced modifications such as deamidation and oxidation
- Chromatographic separation by reversed-phase liquid chromatography (RPLC) coupled with mass spectrometric detection
- Confident identification of peptides, enabling complete sequence coverage of both innovator and biosimilar mAbs
- Relative quantification of peptide post-translational modifications to support biosimilarity assessment
Section Overview
Introduction
Monoclonal antibodies (mAbs) have revolutionized the treatment of a broad spectrum of diseases, ranging from cancers to autoimmune and infectious diseases, by targeting specific antigens with high precision. This specificity allows mAbs to effectively neutralize pathogens and regulate immune responses, thereby offering substantial therapeutic benefits.
Biosimilars, developed as highly similar versions of the original innovator mAbs, offer comparable therapeutic advantages at lower cost. The rising importance of biosimilar development is driven by the need to provide more affordable treatment options as patents for many innovator mAbs expire. During a biosimilar development program, mAbs undergo rigorous analytical and clinical evaluations to meet stringent regulatory standards and to demonstrate similarity in terms of efficacy, safety, and quality. This extensive validation ensures that biosimilars deliver similar clinical outcomes while providing global access to essential therapies.1
Although biosimilars share the same amino acid sequences as the innovator drugs, structural differences may arise due to post-translational modifications (PTMs) influenced by variations in the manufacturing process. These modifications, such as N-glycosylation and changes to terminal residues, present significant challenges in establishing comparability between biosimilars and their reference products.2,3
LC-MS-based peptide mapping is a powerful tool in biosimilarity assessment, as it provides precise and comprehensive analysis of protein structures. This technique enables detailed comparison between a biosimilar and its reference product by generating detailed "fingerprints" of the protein. The resulting data set allows detection of even subtle differences in amino acid sequences and PTMs. Such analysis is crucial, as peptide modifications, including glycosylation or oxidation, can impact protein function, stability, and immunogenicity. By ensuring that the biosimilar closely mirrors the original biologic in structure and function, LC-MS-based peptide mapping plays a key role in confirming biosimilarity and ultimately safeguarding product efficacy and safety. Moreover, to achieve an accurate representation of the therapeutic protein, it is essential to minimize protein degradation and modification during sample preparation.4 In this work, an optimized approach for enzymatic digestion and peptide mapping is presented, using trastuzumab as a model. The method employs filter-assisted sample preparation (FASP) together with a proprietary buffer solution designed to reduce amino acid oxidation and deamidation, thereby enhancing peptide recovery and accuracy.
Trastuzumab, a humanized IgG1 monoclonal antibody that is now off-patent, has demonstrated high efficacy in the treatment of HER2-overexpressing breast cancer. Its mechanism of action involves binding to the Fc gamma receptor III (FcγRIII) in the Fc region, thereby promoting immune cell activity through antibody-dependent cell-mediated cytotoxicity (ADCC). Initially approved in the United States in September 1998 and in the European Union in August 2000, trastuzumab has since been followed by the approval of five biosimilar versions in both regions. 5,6
Experimental
Reagent Preparation
~8 M Urea solution
Dissolve 2.25 g of urea (51456) in 3 mL of water. The solution may be warmed slightly to aid in dissolution while mixing, but the temperature should not exceed 30 °C. Heating above 30 °C promotes the formation of isocyanates. Always use freshly prepared urea solution to avoid isocyanate formation and possible carbamylation of proteins.
100 mM Tris(2-carboxyethyl)phosphine (TCEP) solution
Prepare 100 mM TCEP by diluting 200 μL of 0.5 M TCEP solution (646547) with 800 μL of 8 M urea solution.
100 mM Iodoacetamide (IAM)
Dissolve one vial (A3221) in 606 μL of 8 M urea to prepare a 500 mM stock solution. Further dilute to a 100 mM working solution by combining 200 μL of the 500 mM IAM stock with 800 μL of 8 M urea. IAM degrades quickly in solution, therefore, always use a freshly prepared solution, and discard the unused material.
SOLu-Trypsin
SOLu-Trypsin (EMS0004) is supplied ready for use at a concentration of 1 mg/mL and can be stored at 2-8 °C up to two years.
Glu-C
Reconstitute the 50 µg vial of Glu-C (P6181) with 50 µL of LC-MS water to prepare a 1 µg/µL solution. Use the appropriate volume of Glu-C according to the decided enzyme-to-protein ratio. For example, for a 1:20 enzyme-to-protein ratio, 2.5 µL of 1 µg/µL Glu-C solution shall be added to digest 50 µg of protein. The reconstituted Glu-C should be stored at -20 °C. When stored at –20 °C, the product retains activity for at least one year. Frozen solutions can be thawed and refrozen repeatedly without loss of activity (as per literature7).
Asp-N
Reconstitute the 2 µg vial of Asp-N (P3303) with 50 µL of LC-MS water to prepare a 0.04 µg/µL solution. Use the appropriate volume of Asp-N according to the decided enzyme-to-protein ratio. For example, for a 1:200 enzyme-to-protein ratio, 6.25 µL of 0.04 µg/µL Asp-N should be added to digest 50 µg of protein. The reconstituted Asp-N can be aliquoted and frozen for several weeks (as per literature8).
Low Artifact Digestion Buffer (LADB)
This product (EMS0011) is supplied ready for use and can be stored at 2-8 °C up to two years.
MS QCAL Peptide Mix
MSQC2 is soluble in solutions compatible with proteomics workflows, such as 0.1% formic acid in water or 0.1% trifluoroacetic acid in water. Centrifuge the vial at 10,000 x g to collect the product at the bottom. After addition of liquid to the vial, vortex rigorously. Aliquot the reconstituted MSQC2 peptide solution and store it at -70 °C for up to 6 months. After solubilization in an appropriate buffer, MSQC2 can be directly injected for LC-MS analysis.
Sample Preparation
Commercially available trastuzumab innovator (Herclon, Roche) and biosimilar (CanMab, Biocon) was procured and reconstituted according to the manufacturer’s instructions. In addition, the SILuTMLite SigmaMAb Trastuzumab (MSQC22) Monoclonal Antibody Standard was included as a preparation control. The mAb samples were denatured, reduced, alkylated, and digested as described below for peptide mapping analysis.
Sample preparation was performed in a 30 kDa molecular weight cut-off filter cartridge (Microcon® Centrifugal Filter, MRCF0R030) which simplifies washing and removal of reagents. The filter membrane retains the mAb of interest until completion of the digestion procedure, after which the resulting peptides pass through the filter while the enzyme used for digestion is retained.
Sample preparation steps
- Monoclonal antibody samples equivalent to 50 µg of protein were placed in a microcentrifuge tube. Ideally, at least 50 μg or more protein should be used. The sample solution should not exceed 100 μL. If the volume is less than 20 μL, adjust to 20 μL with water.
Note: Exchange the buffer to 8 M urea if the sample solution is known to interfere with reduction or alkylation
Denaturation & Reduction - Add 100 μL of 100 mM TCEP at a pH of 6.5.
- Incubate for 30 min at 37 °C while mixing at 300 rpm. Transfer the reduced sample to a 30 kDa Microcon® filter (MRCF0R030) in a collection tube.
Note: Do not use Amicon® filter or any type of membrane other than Microcon® nitrocellulose 30K cut-off filter. - Centrifuge at 14,000 x g for 5 minutes*.
- Discard the flow-through from the collection tube
Alkylation - Add 100 μL of 100 mM IAM and vortex for 1 minute.
- Incubate for 20 minutes at ambient temperature in the dark while mixing. Centrifuge at 14,000 x g for 5 minutes.
- Discard flow-through from the collection tube.
- Add 150 μL of LADB to the filter unit and centrifuge at 14,000 x g for 5 minutes*. Repeat this step once more. Transfer the filter unit to new collection tube and add 100 μL of LADB to each filter unit.
Digestion - Add the enzyme in an amount sufficient to achieve the desired enzyme-to-protein ratio for complete digestion.
- Wrap the filter unit with PARAFILM® to prevent evaporation.
- Vortex for 1 minute.
- Incubate the filter unit at 37 °C for 4 hours while mixing at 300 rpm.
- Centrifuge the filter unit at 14,000 x g for 5 minutes*
- Add 40 μL of LADB and centrifuge the filter unit at 14,000 x g for 3 minutes*. Repeat this step once more.
- Add 2 μL of neat formic acid to the filtrate, vortex, and centrifuge briefly. (This step is optional since some peptides may precipitate at low pH).
- Analyze the digested samples immediately or store at –80 °C until LC-MS/MS analysis.
* Note: If the solution has not been fully collected, centrifuge for an additional two minutes. Do not centrifuge longer than necessary, as extended centrifugation may increase oxidation (LADB Technical Bulletin).
Comments on obtaining higher sequence coverage
- Vacuum drying can be used if less than 50 μg of protein is available initially. This step further concentrates the sample.
- Some large and hydrophobic peptides may not pass through the FASP filter. To prevent loss of these peptides and improve sequence coverage after trypsin digestion, collect the solution remaining above the filter and transfer it to a new collection tube. Centrifuge the filter unit at 14,000 x g for 5 minutes. Add 40 μL of LADB and centrifuge the filter unit again at 14,000 x g for 3 minutes. Repeat this step once more. Combine all solutions, including both filtered and unfiltered.
LC-MS/MS System Set-up
Instrument Calibration
The Waters QToF Xevo G2XS mass spectrometer was calibrated over a mass range of 50 – 2500 m/z using a 10 μL/min infusion of 0.4 mg/mL sodium iodide in IPA:water (1:1).7
System Suitability
SILuTMLite SigmaMAb Trastuzumab (MSQC22) Monoclonal Antibody Standard and MS QCAL Peptide Mix (MSQC2) were evaluated to verify sample preparation quality and system performance. Test samples were bracketed with system suitability samples to confirm the validity of test sample results. (The sequence of MSQC22 is shown in Appendix).
Data Analysis
The obtained data was processed using the peptide mapping tool in Waters_Connect software to identify peptides using the MaxEnt3 algorithm and the trastuzumab sequence (digested with specific protease). The amino acid sequence of trastuzumab is provided in the Appendix.
The targeted PTMs included one fixed modification, carbamidomethylation (C), and the following variable modifications: pyroglutamic acid formation at the N-terminus of glutamic acid (E), C-terminal lysine truncation, oxidation of methionine (M) and tryptophan (W), deamidation of asparagine (N) and glutamine (Q), and glycosylation at asparagine (N) with glycoforms G0F, G1F, and G2F.
The data was filtered to report only peptides with mass tolerance of ± 10 ppm at both MS and MS/MS level. Furthermore, peptide identifications for each enzyme were confirmed based on strong MS response, consistent MS/MS fragmentation pattern, and low mass error.
Results and Discussion
System Suitability
MS QCAL Peptide Mix
This standard (MSQC2) consists of 22 peptides derived from tryptic digestion of the QCAL protein concatemer, with peptide masses ranging from 415 to 3,100 Da. Figure 1 displays the integrated base peak intensity (BPI) chromatogram of MS QCAL Peptide Mix. The identified peptides and their molecular masses matched the theoretical masses, listed in the product information sheet.
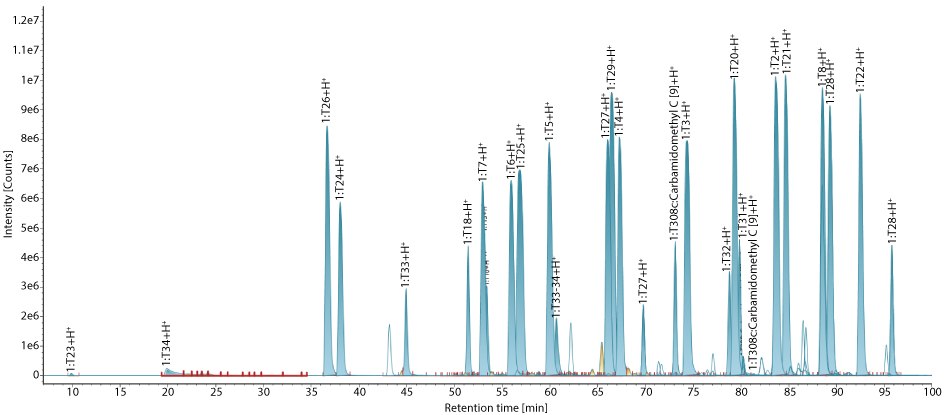
Figure 1.Base peak intensity (BPI) chromatogram of MS QCAL Peptide Mix. All peptides of MS QCAL are identified with good MS response, fragmentation, and low mass error.
MS QCAL Peptide Mix is employed as a system suitability standard to verify MS system performance, offering multiple components for evaluation. This standard is injected at the beginning, middle, and end of the analytical sequence. A critical performance metric is the ability to distinguish between two peptides with a mass difference of 0.0364 Da. In MS QC Cal, peptides 1:T2 and 1:T8 are nearly identical, differing only by a Lys→Gln substitution, which results in this small mass variation. The ability to resolve these two peptides, and achieve confident identification, serves as a benchmark for assessing instrument resolution. Table 3 shows that these two peptides were successfully resolved and detected with high mass accuracy within 1.0 ppm.
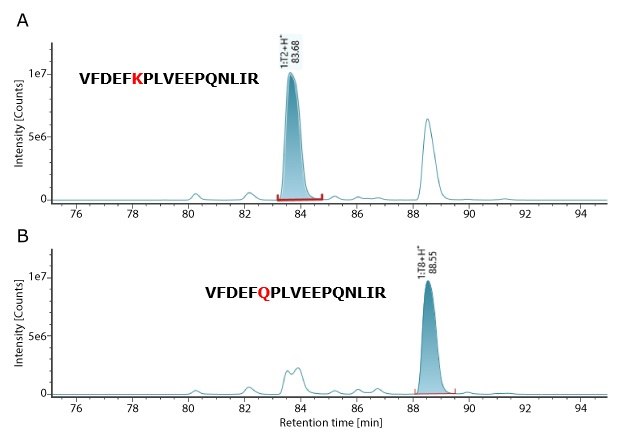
Figure 2.XIC Chromatogram of MS QC Cal peptides-1:T2 and 1:T8.
SILuTMLite SigmaMAb Trastuzumab Monoclonal Antibody Standard
This monoclonal antibody standard (MSQC22) was assessed as a control for sequence coverage. The chromatographic analysis, shown in Figure 3, provided a high coverage of 97% (Figure 4) indicating successful sample preparation and confirming that the system was functioning as intended.
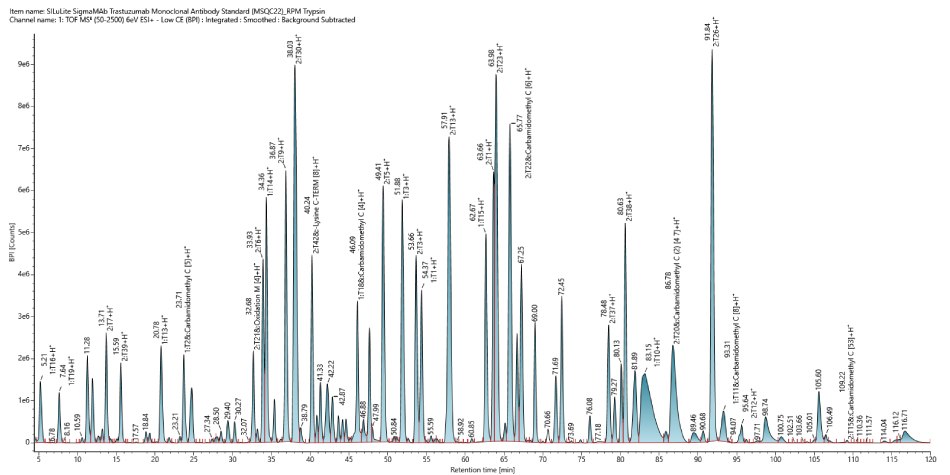
Figure 3.Base peak intensity (BPI) chromatogram of SigmaMAb Trastuzumab Monoclonal Antibody Standard MSQC22).
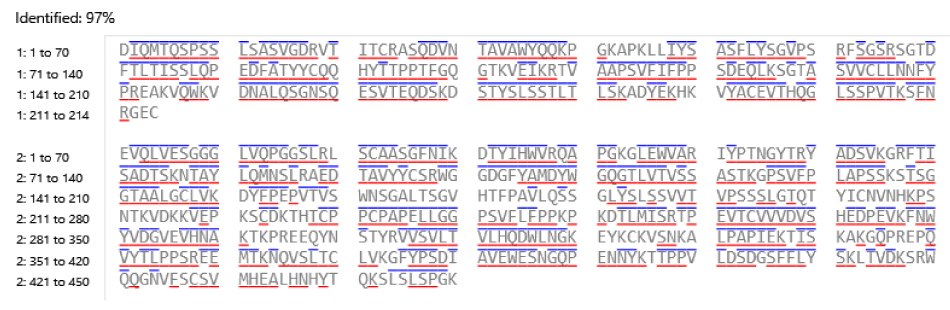
Figure 4.Sequence coverage of SigmaMAb Trastuzumab Monoclonal Antibody digested using trypsin. Amino acids within each confirmed peptide with a blue line above or red line below indicate detection of their respective b and y ions.
Samples
For biosimilarity assessment, peptide maps obtained from the LC-MS/MS analysis were compared using the binary compare mode within the UNIFI module in Waters connect platform. Figure 5 displays mirror plots of the BPI chromatograms of the innovator and biosimilar mAbs. Sequence coverage of 95% or higher was achieved for both the innovator and biosimilar using separate trypsin and Asp-N digestions, while Glu-C digests provided 65% sequence coverage. When combined, sequence coverage reached 100% with high mass accuracy (within 15 ppm) and supporting fragment ions in the high energy data channel (b/y ions ≥ 3). The Ascentis® Express Peptide ES-C18 column provided efficient peptide separation, enabling accurate peptide identification. Paired with the analytical sensitivity and precision of advanced mass spectrometry, this column supports effective comparison of innovator and biosimilar molecules and allows differentiation between closely related sequences in complex mixtures. Figure 6 shows the sequence coverage maps of the innovator and biosimilar (light and heavy chains) for each enzyme digest. Missing peptides, primarily tripeptides, were not detected due to their low retention on the selected stationary phase under the applied chromatographic conditions, leading to early elution and limited MS detectability.
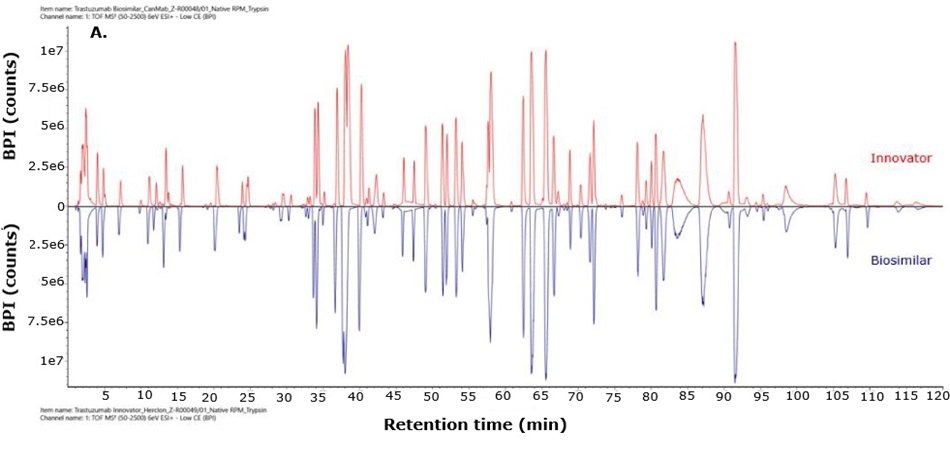
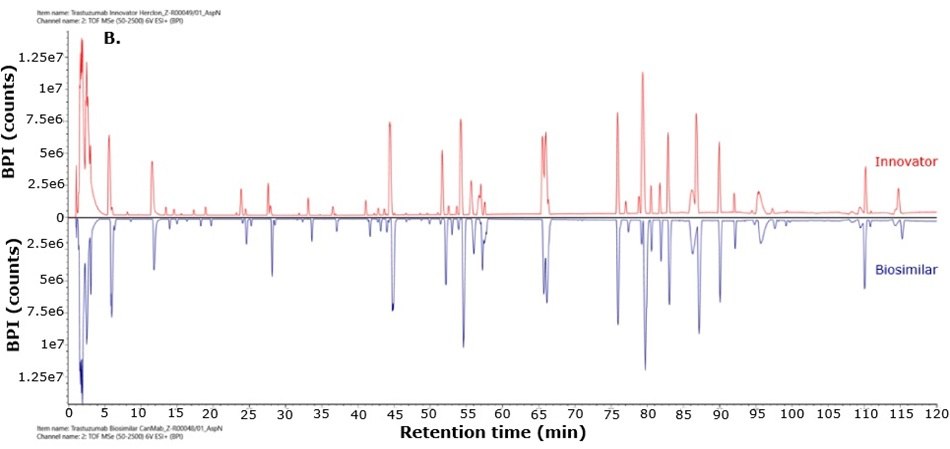
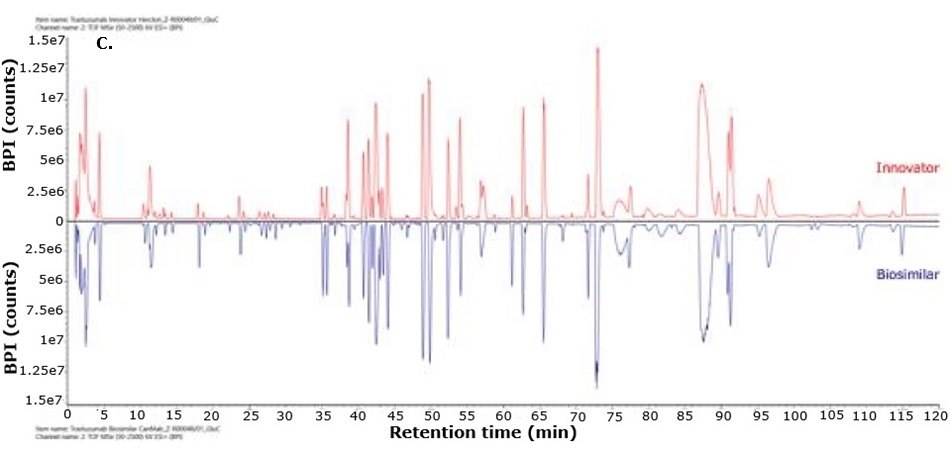
Figure 5.Mirror plot of BPI chromatograms of trastuzumab innovator (red) and biosimilar (blue) digested with trypsin (A), Asp-N (B) and Glu-C (C).

Figure 6.Sequence coverage map of trastuzumab innovator and biosimilar digested with trypsin (A), Asp-N (B) and Glu-C (C). Amino acids within each confirmed peptide having blue line above or red line below indicates the detection of their respective b and y ions.
Although the chromatograms appeared similar, minor differences in the relative abundance of modifications were observed. Site-specific PTMs were characterized using MS/MS spectra acquired under high collision energy. The resulting fragmentation patterns, showing high coverage and matching of fragment ions across each peptide's amino acid backbone, enabled accurate peptide identification and localization of modification sites at specific amino acids within the peptide sequence. N-glycosylation modifications, pyroglutamation, and lysine truncation were evaluated in both the innovator and biosimilar mAbs for biocomparability. The well-conserved heavy chain DLTMISR and heavy chain GFYPSDIAVEWESNGQPENNYK sequences were used as markers to assess the extent of oxidation and deamidation, respectively. High energy spectra (MSE) for a heavy chain tryptic peptide, 2:T1 (DTLMISR), in its native and oxidized forms are shown in Figure 7, confirming the site of modification. The relative extent of modification on specific peptides was quantified based on extracted ion chromatogram (XIC) response of native and modified peptides, in both innovator and biosimilar mAbs. Histogram comparisons, illustrated in Figure 8, provide a quick means of tracking and evaluating quality attribute levels across innovator and biosimilar samples.
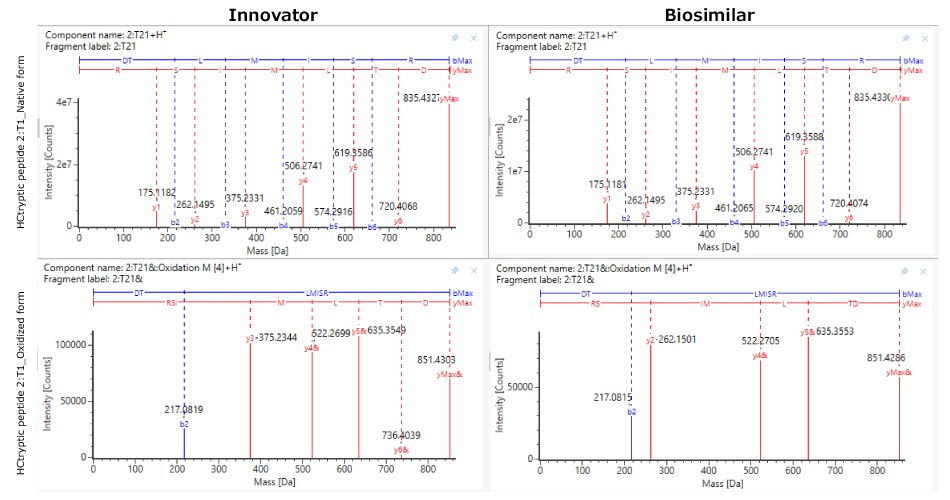
Figure 7.Representative LC-MSE high energy fragmentation spectra of heavy chain tryptic peptide 1:T1 in its native (top) and oxidized form (below) in innovator (left) and biosimilar (right).
The minor differences observed are acceptable within the regulatory framework1, as they do not have meaningful impact on clinical efficacy or safety. The key criterion being that any molecular differences must not impact the safety, efficacy, or immunogenicity profiles of the biosimilar.
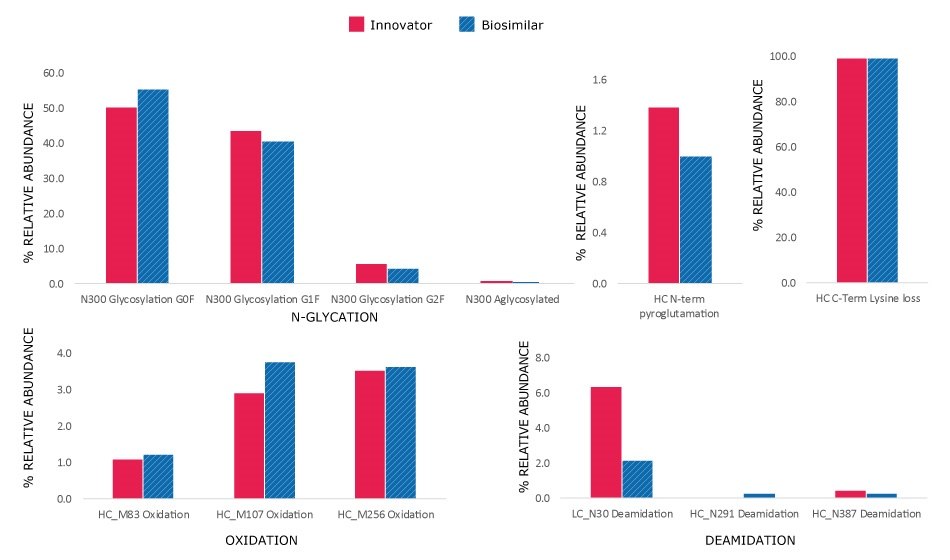
Figure 8.Histograms showing relative abundance of modifications in innovator and biosimilar trastuzumab.
Conclusion
Comprehensive characterization of biosimilar monoclonal antibodies (mAbs) is essential to ensure their safety and efficacy, leveraging the established performance of the innovator product to mitigate potential clinical challenges associated with the follow-on product. This application note outlines a peptide mapping method using reversed-phase liquid chromatography-mass spectrometry (RPLC-MS) with an Ascentis® Express ES-Peptide C18 column, providing the necessary resolution required to efficiently characterize monoclonal antibodies. The workflow was developed for peptide mass analysis of reduced monoclonal antibodies, using trastuzumab as a model mAb and a SILuTMLite SigmaMAb product as a preparation control. The study focussed on assessing the comparability of the innovator drug and its biosimilar counterpart. The method includes digestion using filter-assisted sample preparation (FASP), followed by data acquisition and processing steps including extraction, sequence matching, sequence coverage computation, PTM identification, and relative quantification.
The low artifact digestion buffer (LADB) is specifically formulated to reduce deamidation and oxidation during the digestion process while maintaining high digestion efficiency. The buffer’s pH is optimized for enzymatic activity and includes a proprietary antioxidant to further protect against oxidation. FASP helped in maximizing recovery of peptides to result in reliable peptide identifications in both native and modified forms. Trastuzumab samples (innovator and biosimilar) yielded 100% combined sequence coverage using trypsin, Asp-N and Glu-C. Identification and quantification of critical peptide modifications highlights the multi-attribute aspect of peptide mapping using this method. Overall, the results demonstrate the effectiveness of the workflow in assessing sequence coverage and critical PTMs between the innovator and biosimilar mAbs, ensuring that the biosimilar's PTM profile aligns closely with that of the innovator product.
Related Products
Samples and System Suitability Reagents
HPLC Column and Sample Preparation
Enzymes
Solvents and Reagents
Accessories
References
Appendix
Structure and Sequence of Trastuzumab
SigmaMAb Trastuzumab Heavy Chain
EVQLVESGGGLVQPGGSLRLSCAASGFNIKDTYIHWVRQAPGKGLEWVARIYPTNGYTRYADSVKGRFTISADTSKNTAYLQMNSLRAEDTAVYYCSRWGGDGFYAMDYWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK
SigmaMAb Trastuzumab Light Chain
DIQMTQSPSSLSASVGDRVTITCRASQDVNTAVAWYQQKPGKAPKLLIYSASFLYSGVPSRFSGSRSGTDFTLTISSLQPEDFATYYCQQHYTTPPTFGQGTKVEIKRTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC
如要继续阅读,请登录或创建帐户。
暂无帐户?