Whole Transcriptome Amplification of RNA from Low Cell-Number Samples
Abstract
The efficacy of amplification of small quantities of total RNA with the Complete Whole Transcriptome Amplification Kit (WTA2) was examined in this study. Total RNA extracted from decreasing numbers of FACS-isolated bone marrow stem cells (10-, 100-, and 1000-cells samples) was amplified with the WTA2 kit. A call rate of 58.8% of unique biological array features was observed for the 10-cell vs. 100-cell microarray analysis, with a similar call rate of 61.46% for 10-cells vs. 1000-cells. Greater than 90% commonality existed between the intersecting data sets for the two analyses. After a more stringent screening (p < 0.0001), the 10-cell vs. 100-cell comparison revealed 5568 intersecting features. The comparable analysis for 10 cells vs. 1000 cells resulted in 4977 features, with 3862 features common to both comparisons. In addition, the effect of decreasing RNA input on amplification efficiency was examined. Results indicated that an adjustment to the library synthesis primer concentration allowed for maintenance of linear amplification and representative qPCR at low RNA input quantities. This adjustment proved to be critical for downstream qPCR applications, but not for the case where the amplification product was used as microarray target. This study confirms that the Complete Whole Transcriptome Amplification Kit can effectively amplify low input quantities of RNA, approaching the single cell level.
Introduction
There is increasing interest in profiling gene expression in samples of decreasingly smaller numbers of cells. To date, RT-PCR remains the gold standard for simultaneously measuring expression of a limited number of transcripts2, 3. Sensitive global measurement of transcript levels can be obtained by exponential (PCR-based) amplification4-8, or in combination with linear methods7. The Complete WTA2 kit, developed for enhanced amplication of damaged RNA (described in an accompanying poster presentation, “Detection of Prostate Cancer Biomarkers by Expression Microarray Analysis of WTA2-Amplified FFPE Tissue RNA”), has been shown to efficiently amplify low input quantitiies of RNA, approaching the single cell level. RNA isolated from as few as 10 fluorescence-activated sorted cells (FACS) is amplified, maintaining expression patterns similar to those observed for 10- and 100-fold larger cell samples. Challenges unique to exponential amplification of small RNA input quantities are discussed and effectively addressed for downstream qPCR and microarray applications.
Materials and Methods
FACS-isolated bone marrow stem cells were provided by Albert Donnenberg, University of Pittsburgh. Cells were sorted directly into GenElute Mammalian RNA Extraction Kit Lysis Buffer (RTN70, Component No. L8285), frozen and stored at -70 °C until extraction. Total RNA was extracted from 10-, 100-, and 1000-cell samples using the GenElute Mammalian RNA Isolation kit. Sample RNA was eluted at a reduced volume of 40 μL with nuclease-free water, and subjected to a scaled-up (5X) RNase-free DNase digestion (AMP-D1), for a final volume of 97 μL. Each RNA sample was amplified in its entirety using the Complete WTA2 kit (Figure 1): the entire heat-attenuated DNase digestion was directly added to the recommended bulk amplification reaction volume of 375 μL, compensating by subtraction of water volume. As per kit instructions, the 375-μL volume was split into five 75-μL reactions for amplification. Each reaction was carried through two cycles beyond stationary amplification (“plateau”, Figure 2). Unincorporated primers and other reaction components were removed from the amplification product using the GenElute PCR Cleanup kit (NA1020). Quantitative PCR (Figure 3A, B) was performed using SYBR® Green JumpStart™ Taq ReadyMix™ (S4438), with 250 nM primer concentration. Primer pairs, with the exception of the 18S rRNA set, are specific for cDNA detection.
Amplified RNA (double-stranded cDNA) was labeled with cy3 or cy5 following the Agilent Genomic DNA Labeling Kit Plus procedure (5188-5309). Two amplified, differentially-labeled cDNA targets were combined, heat-denatured, applied to Agilent Whole Genome™ arrays (G4112F), and incubated at 65˚ C for 40 hours. Hybridization and subsequent wash procedures for the Agilent Oligonucleotide Array-Based CGH for Genomic DNA Analysis were followed, except for the omission of COT-1 DNA block during hybridization. Arrays were scanned (Agilent scanner model G2505B) and features extracted. Intensities representing non-uniform features or associated with local non-uniform background, in addition to “absent” and saturated array feature intensities were removed using Microsoft Excel™. All intensities less than 100 were removed, except where indicated. Intensities were then subjected to median normalization, Welch’s t-test, Benjamimi-Hochberg false discovery rate correction, and a ± 1.5-fold differential threshold using Genesifter pair-wise comparison, except where only median normalization was performed, as indicated. Normalized-only intensities and intensities subjected to Genesifter™ (Geospiza) pair-wise analysis were further analyzed using the Genesifter Intersector software and Spotfire™ (TIBCO), where indicated.
For the study of the effect of reduced library synthesis primer concentration on amplification of decreasing quantities of RNA input, mRNA levels for high- and mediumcopy transcripts in the amplification products, plus a “problematic” low-copy transcript, were evaluated by real-time qPCR. Preparation of the lowered concentrations of library synthesis primers was accomplished by dilution of the kit Library Synthesis Solution (component no. L9293) with 5 mM dNTPs (a 2-fold aqueous dilution of 10 mM dNTPs, (D7295).
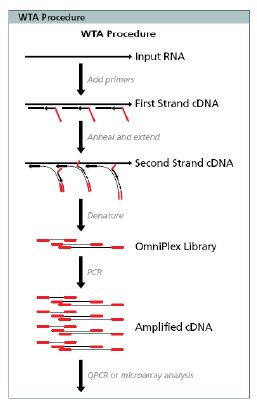
Figure 1. WTA Methodology1. Total RNA is combined with Library Synthesis Solution and heat-denatured. This is followed with the addition of Reaction Buffer and the strand-displacing Library Synthesis Enzyme, for “single-tube”reverse-transcription and second-strand cDNA synthesis (Omniplex™ Library). The 3’ end of the Library Synthesis Primer is “quasi-random” and substantially non-self-complementary, while the 5’ end is a single, constant non-self-complementary sequence that serves as the annealing site for the universal amplification primer.
Results and Discussion
Amplification of Cellular RNA. Total RNA was extracted from FACS-isolated from human bone marrow stem cells as described in the Materials and Methods. Amplification for all samples was allowed to proceed through 2 cycles of “plateau”, or stationary amplification (Figures 1 and 2). Amplification product was evaluated by qPCR (Figure 3A, B) and agarose gel electrophoresis prior to microarray analysis (Figure 3C).
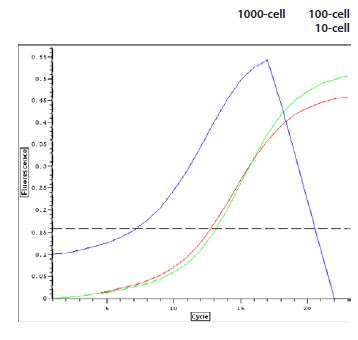
Figure 2. RNA Amplification Profiles. Total RNA was extracted from cell samples as described in Materials and Methods. RNA amplification was allowed to proceed through 2 cycles of stationary amplification (observed by real time PCR using SybrGreen dye). One thousand-cell reactions were stopped at 19 cycles; 100- and 10-cell samples at 23 cycles.
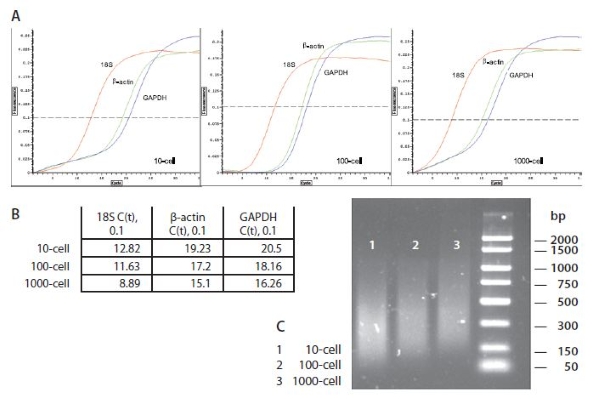
Figure 3. Amplification Qualitative Test. Prior to microarray analysis, amplification reaction products were evaluated by real-time qPCR using primer pairs for 18S, GAPDH, and β-actin transcripts (A, B). qPCR amplicons were further checked by 1% agarose electrophoresis (C).
Microarray Analysis of Whole Cell Samples. A strong concordance between the three stem cell samples is observed for detection of microarray features, in this case, normalized intensities (> 100) (Figure 4). Greater than 85% of features detected in the 10-cell sample are also detected for the 100-cell sample, while > 90% of 10-cell features are also detected in the 1000-cell sample. Taking into account ing a total number of 41,000 unique biological probles on the Agilent 4 x 4 array, detected features for the 10-cell and 100-cell comparison comprised 59% of total array features, with 48% of array features shared between the data sets. Similarly, for the 10-cell and 1000-cell sample comparison, 61% of total array features are detected, with 52% of features shared .
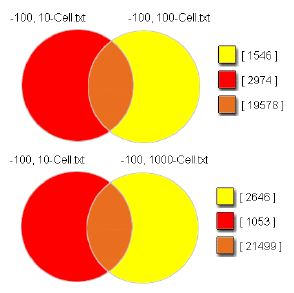
Array Features Detected
Median-Normalized
Intensities > 100
Figure 4. Microarray analysis revealed a strong concordance between the cell samples. Overlapping regions represent median-normalized intensities (> 100). Greater than 85% of detected features represented in the 10-cell sample are detected for the 100-cell sample, while > 90% of 10-cell features are also detected in the 1000-cell sample. In addition, the overlapping regions are highly concordant, with greater than 98% of 10-cell/100-cell overlapping intensities detected in the 10-cell/1000-cell intersection.
Application of statistical tests provides additional confidence to the normalized data (Figure 5). An excess of fifty-five thousand features were common for the 10-cell and 100-cell samples at a p < 0.0001. (This stringency reflects the analysis of an n = 4 for each data point.) Moreover, 70% of features common to the two data sets (the 10-cell/100-cell comparison) were also detected in the 10-cell/1000-cell intersection. It is hypothesized that the WTA2 amplification method is sufficiently sensitive to detect basal expression. Such highly stochastic signal(9), however, is eliminated with replication and stringent statictical testing.
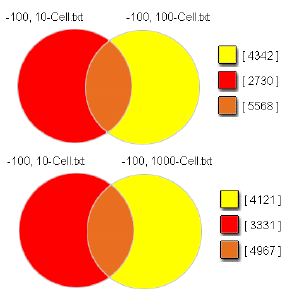
Array Features Detected
Median-Normalized
Intensities >100
p<0.0001
Figure 5. A substantial number of statistically significant microarray features (> 5500) are detected, with concordance between the two overlapping regions of the cell sample data sets maintained following stringent statistical analysis and corrections of the data presented in Figure 3 (see Materials and Methods). Seventy percent of 10-cell/100-cell overlapping intensities are detected in the 10-cell/1000-cell intersection.
Effect of Library Synthsis Primer Concentration on Downstream qPCR Application. It was hypothesized that the relatively high LS primer concentration may lead to overpriming, resulting in shortened amplification product difficult to detect by qPCR. To test this idea, decreasing human prostate RNA input quantities were matched with similarly decreasing LS primer concentration during the library synthesis step. Serial dilutions of RNA input ranged from 25 ng to 12.2 pg, and LS concentration, from the 1X kit concentration to 0.0078X. A marked loss in linearity of amplification efficiency is observed when using the 1X LS primer concentration at RNA input quantities less than ~800 pg (Figure 6A). Linearity in efficiency returns for lower RNA input quantities as the concentration of the 1X LS primer concentration is reduced by 2-fold increments (Figure 6B-D). Quantitative PCR results using primer sets for high- (18S rRNA), medium- (beta-actin, GAPDH), and low-copy TMCC2 transcripts corroborate these observations (Figures 7A-D). Interestingly, TMCC2, indicating only a background signal for an intermediated RNA input quantity, at the 1X LS primer concentration, while amplification product is detected for all RNA input quantities at an 8-fold LS primer dilution, suggesting an 8-fold dilution of the WTA2 LS primer is optimal for TMCC2 for downstream qPCR.
Figure 6. The effect of library synthesis primer concentration on amplification of small quantities of input RNA was examined. Two-fold serial dilutions of human prostate total RNA provided for reaction input quantities ranging from 25 ng to 12.2 pg. In addition, decreasing concentrations of library synthesis primer were evaluated: the 1X WTA2 kit concentration and three additional 2-foldfold serial dilutions. C(t) values measured at early exponential growth (threshold, 0.025) for each amplification reaction are plotted against the input quantity of RNA for each library synthesis primer concentration tested. The results for the kit concentration primer (1X) and a 2-, 4-, and 8-fold dilution of primer are shown. These results suggest that RNA input quantities less than ~800 pg. are not efficiently amplified in the presence of 1X library synthesis primer concentration. Intermediate primer concentrations (0.5X and 0.25X) show increasing improvement of amplification efficiency. Overall efficiency appears to be maintained when the 8-fold dilution of library synthesis primer is used.
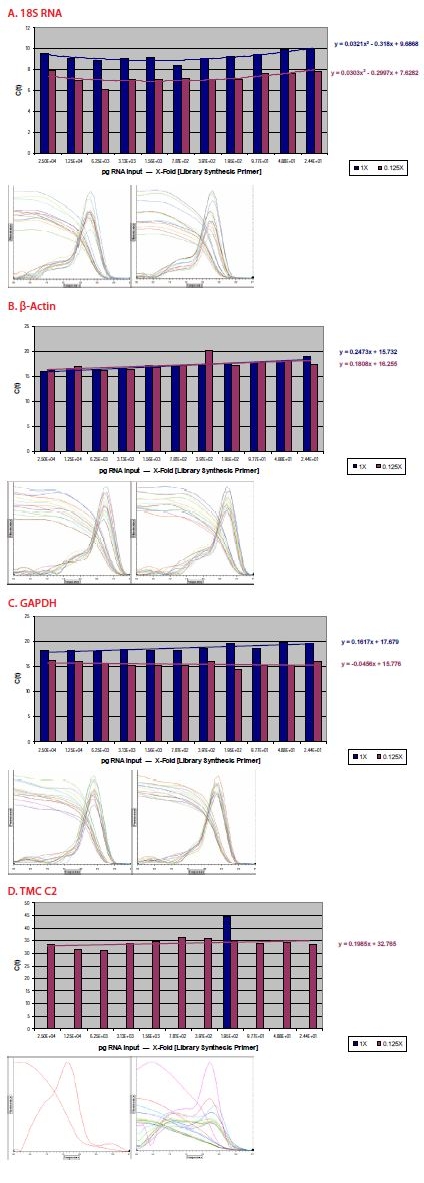
Figure 7. Amplification products were subjected to qPCR evaluation using primer sets for 18S rRNA (A), β-actin (B), GAPDH (C), and TMCC2 (D). Inserts show the melt curves for amplification product at the decreasing RNA reaction input quantities, and for the 1X and 0.125X primer concentrations. A – C represent high-copy rRNA and medium-copy and house-keeping transcripts; TMCC2, human transmembrane and coiled-coil domain family 2 transcript, was noted to be inconsistently detectable in previous studies, with C(t)s in the mid-30s. Interestingly, a pattern of biphasic inefficient amplification is observed for 18S rRNA, at highest and lowest input quantities for both library synthesis primer concentrations. Generally consistent C(t) values are observed for β-actin and GAPDH at the 0.125X primer concentration, while, a gradual decrease in efficiency is observed as input quantities decrease at the 1X primer concentration. At the 1X primer concentration, no amplification product is detected for TMCC2. Only a spurious signal at 75 degrees (primer-dimer, gel data not shown) is observed. However, at the low library synthesis prime concentration, TMCC2 amplicon is detectable at all input quantities.
Effect of Library Synthsis Primer Concentration on Downstream Microarray Analyses
To determine whether the use of a reduced LS primer concentration is appropriate for microarray anaysis of low RNA input quantities, amplification product from three input quantities was tested (Figure 8): 6.25 ng (approximately the kit-recommended quantitiy per a 75-μL reaction)1, 391 pg (just under the 800-pg demarcation observed in Figure 6A)2, and 48.8 pg (the approximate mass equivalent for a ‘single’ cell).3 Amplification products were labeled and subjected to microarry analyses as described in Materials and Methods. In Figure 8A (non-normalized intensities) reproducibility between same-dye duplicates for the input quantities tested is shown. Figure 8B (also non-normalized intensities) shows a crosscomparisons of input duplicates (n = 4). A predictable increase in variance is observed for each comparison for the 1X LS primer concentration (see discussion). However, there is considerably less quantitative and qualitative correspondence between the duplicates generated with the 8-fold dilution of LS primer. This observation extends to Figure 8C, where the data sets have been statistically analyzed (t-test, false discovery rate correction, and ± 1.5 threshold). The 1X library synthesis primer concentration performs indisputably better for microarray analysis.
Figure 8. Amplification product generated from 6.25-ng, 390-pg, or 48-pg input quantities, prepared with 1X or 0.125X library synthesis primer, was labeled for microarray analysis as described in Materials and Methods. Results are shown in Panels A through C (log intensities vs. log2 intensities). Panel A shows reproducibility for duplicate 6.25-ng reactions (1), 390-pg reactions (2), and 50-pg reactions (3). In Panel B, 6.25-ng reaction intensities are plotted versus 390-pg reaction intensities (1), 6.25-ng reaction intensities versus 48-pg reaction intensities (2), and 390-pg reaction intensities are plotted versus 48-pg reaction intensities (3). Statistically analyzed and normalized intensities are shown in Panel C: 6.25-ng reaction intensities versus 390-pg reaction intensities (1), 6.25-ng reaction intensities versus 48-pg reaction intensities (2), and 390-pg reaction intensities versus 48-pg reaction intensities (3). The positive effect of lowered library synthesis primer concentration observed for quantitative PCR is not evident in microarray results.
Conclusions
These observations indicate that reducing the library synthesis primer concentration enhances detection of low-copy transcripts when performing downsteam qPCR
application. At 1X LS primer concentration, medium-copy transcripts can be reliably detected (Figure 7B, C). However, low-copy transcripts may be over-primed, resulting in template too short for both qPCR primers of a set to find their respective annealing sites (Figure 7D). This does not appear be a problem for microarray analysis (Figure 8), where size reduction of the amplification product would be essentially equivalent to target fragmentation. However, reduced primer concentration decreases the probability of the LS primer to finding all potential templates, thus preventing uniform amplification of the transcriptome. This is not to suggest that the results of microarray analysis are not affected by higher LS primer concentration. This effect manifests itself in predictably small increases in expression differentials for low-copy transcripts and a similarly minor compression of differentials associated with high-copy transcripts. Both are manageble by simple scaling algorithms10. Therefore, the reduced library synthesis primer concentration is optimal for detection of low-copy transcripts by qPCR, and the standard kit concentration is used for representatative amplification for all RNA input quantities when following with microarray analysis.
(For the investigator utilizing WTA2 amplification product for eventual qPCR studies, it is suggested that serial dilutions of the LS primer are tested to find the concentration optimal for detection of selected transcripts, as demonstrated in this presentation. Directions for LS primer dilution are provided in the Materials and Methods.)
In conclusion, the Complete WTA2 kit provides a robust, sensitive method for amplifying RNA quantities approaching the single-cell level, while maintaining the representative pattern of expression of higher quantities of RNA input.
Acknowledgements:
Our appreciation to Albert Donnenberg, University of Pittsburgh for providing bone marrow stem cell for this study.
Materials
References
如要继续阅读,请登录或创建帐户。
暂无帐户?