Auto2D® Automated Gel Electrophoresis Device for Fast and Reproducible 2D Gel Electrophoresis
Curious how you can do 2D gel electrophoresis in under 2 hours? Read on to learn about the Auto2D® gel electrophoresis device that offers simple and quick automated gel electrophoresis with higher reproducibility. It can be used in applications such as plant sample analysis, food allergy research, anti-HCP antibody validation, antigen profiling, and more.
Section Overview
- How Does 2-D Gel Electrophoresis Work?
- Fully Automated 2-D Gel Electrophoresis
- How Do You Run a 2-D Gel Using the Auto2D® 2-D Electrophoresis Device?
- Why Do We Use 2-D Gel Electrophoresis?
- Auto2D® 2-D Gel Electrophoresis for Food Allergy Applications
- Simple Automated Gel Electrophoresis for Analyzing Plant Tissue-Specific Proteomes
- How To Obtain More Resolution When Separating Target Proteins In Protein Expression and Purification with Automated 2D Electrophoresis
- Auto2D® System for Anti-HCP Antibody Validation for ELISA and Other Immunoassays
- Auto2D® System for Antigen Profiling
- Auto2D® System for Regulatory Compliance
- TotalLab Software Enables Enhanced 2-DE
- Related Products
How Does 2-D Gel Electrophoresis Work?
Two-dimensional gel electrophoresis (2-DE) gives a higher resolution of protein separation by separating proteins by their isoelectric point (pI) and by their molecular weight.
What is the Difference Between 1D and 2D Gel Electrophoresis?
One-dimensional protein gel electrophoresis techniques such as SDS-PAGE are routinely performed in the lab and the proteins are separated by molecular weight. Whereas 2-DE separates proteins in complex samples by pI value and molecular weight, enabling the direct comparison of hundreds or thousands of proteins simultaneously with high resolution. When paired with analytical software, immunodetection, or mass spectrometry techniques, 2-DE provides a powerful tool that aids in protein identification and other proteomic analyses.
For additional resources and products, please visit our Protein Electrophoresis and Western Blotting page.
Fully Automated 2-D Gel Electrophoresis
Two-dimensional gel electrophoresis has generally been regarded as difficult to perform and time-consuming, requiring advanced user training while offering low reproducibility and high inter-operator variability. The Auto2D® system fully automates two-dimensional gel electrophoresis. The applied sample enters the first-dimension gel passively, separating proteins by their isoelectric points. After equilibration, the first-dimension gel is set onto a horizontal SDS-PAGE gel, which resolves proteins by molecular weight. The Auto2D® 2-D Electrophoresis Device is user-friendly and enables two-dimensional electrophoresis in 1-2 hours for faster, more reproducible results. Additional features and benefits of the Auto2D® 2-D Electrophoresis Device include:
- Easy to use, with no advanced training required
- Quick workflow implementation for reduced lab downtime
- Decreased inter-operator variability and higher reproducibility
- IQ/OQ support for GMP compliance
How Do You Run a 2-D Gel Using the Auto2D® 2-D Electrophoresis Device?
The Auto2D® 2-D Electrophoresis Device fully automates two-dimensional gel electrophoresis, simplifying protein analysis and providing more consistent, reproducible results that are user-independent. The efficient engineering of the Auto2D® system significantly reduces the amount of time spent during sample loading, isoelectric focusing, equilibration, and SDS-PAGE from 4-24 hours to only 1-2 hours. This makes the Auto2D® device unique compared to other semi-automated 2-DE systems on the market.

Figure 1.Two-dimensional gel electrophoresis workflow and the steps that are automated by the Auto2D® device.
Why Do We Use 2-D Gel Electrophoresis?
The Auto2D® 2-D Electrophoresis Device has been proven in numerous applications. Examples of suitable use cases include differential protein expression analysis in cancer research and the elucidation of disease mechanisms. Additional applications include the use for separating purified proteins for crystallization or post-translational modification analysis and 2-D western blotting in research areas such as allergy research or cell signaling pathway analysis.
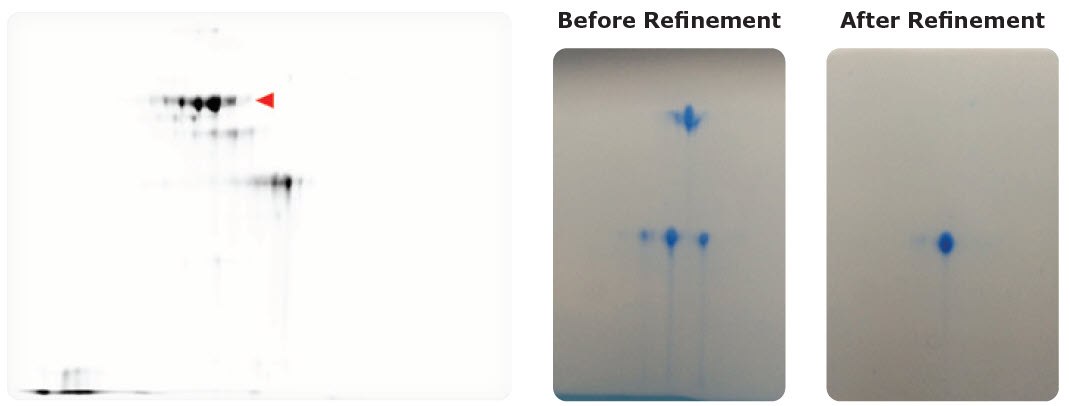
Figure 3.Left: Assessing microheterogeneities in therapeutic antibody drug products. Red triangle indicates spots for target antibody; lower spots are antibody missing regions. Right: Protein analysis before and after purification. 2-DE can be used to check whether the sample is homogeneous for crystallization and X-ray structural analysis.
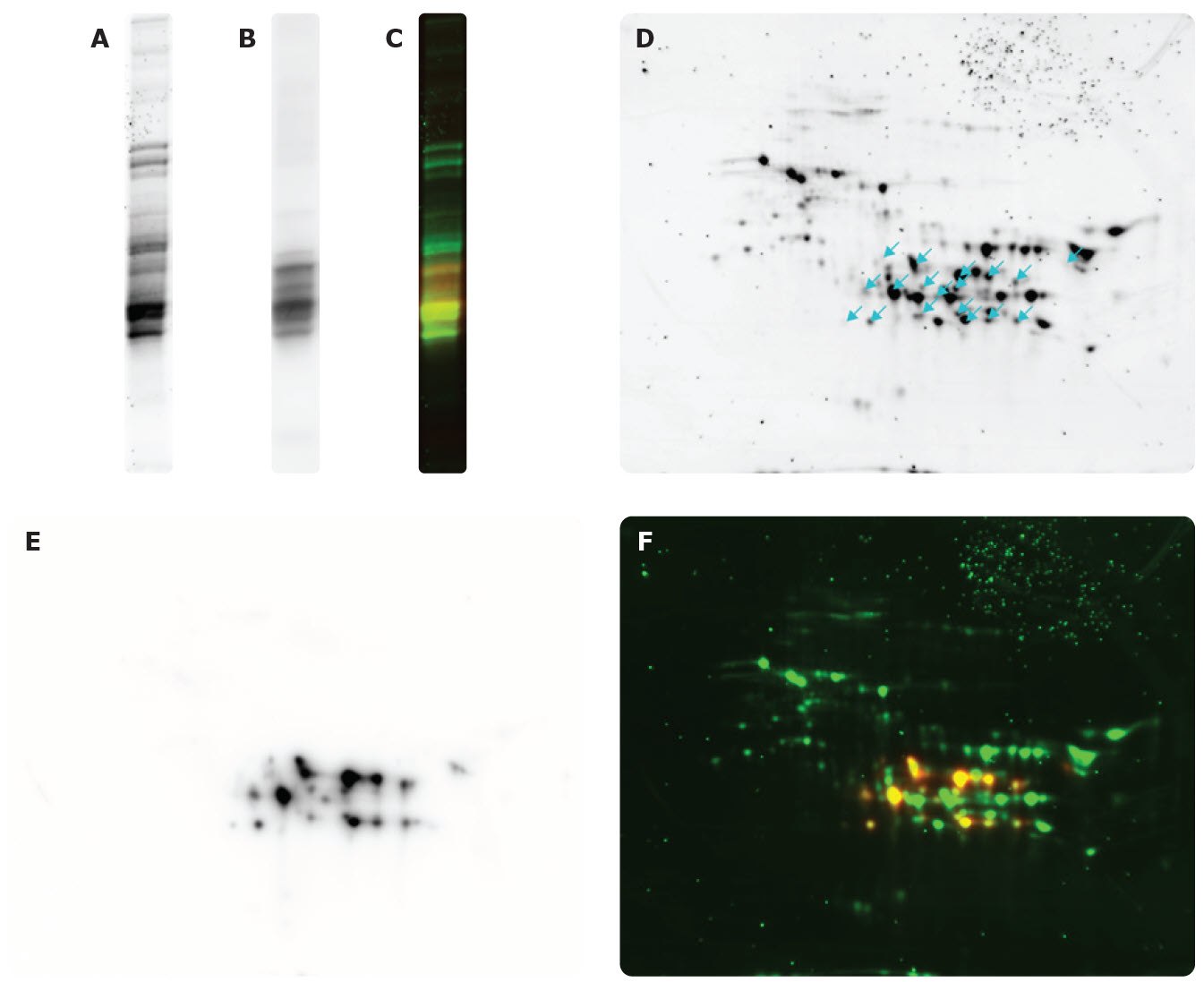
Figure 4.Flour extract proteins (detected by SYPRO Ruby) by SDS-PAGE (A), WB: Anti-Gliadin antibody by SDS-PAGE (B), and merge by SDS-PAGE (C). Flour extract proteins (detected by SYPRO Ruby), arrows indicate spots detected by Anti-Gliadin antibody (D). WB using Anti-Gliadin antibody (E), and merge (F).
Auto2D® 2-D Gel Electrophoresis for Food Allergy Applications
Given the large and growing number of people who suffer from allergies worldwide, it is increasingly important to detect and identify allergens, particularly in food sources. Two-dimensional gel electrophoresis (2D-E) is traditionally used for high-resolution protein separation necessary for the immunodetection of allergenic proteins. However, this traditional separation process is often time-consuming and technically challenging. Our Auto2D® system enables fully automated 2-D electrophoresis in 1-2 hours with quick and reliable results. Here we show that our Auto2D® system is suitable for identifying allergenic proteins from multiple different sources.
Allergies are a common worldwide health issue, with the three major sources of allergens including food (estimated to affect 4% of the world population), pharmaceutical drugs (estimated 10% of the population), and plant pollen (estimated 10-30% of the population and rising).1-3 Allergic reactions occur through an IgE-mediated immunological response and have traditionally been tested through a skin prick test; however, allergen testing, and diagnostics can also be achieved through molecular approaches.1 The radioallergosorbent test (RAST) and the enzyme-linked immunosorbent assay (ELISA) are two methods used to detect and quantify IgE against specific allergens. Since RAST and ELISA do not include a protein separation step, cross-reactivity between antibodies and food matrix components can cause false-positive results in some instances.1
Gel electrophoresis is a common and widely used protein separation method. While standard one-dimensional gel electrophoresis separates proteins based on their molecular weight, two-dimensional gel electrophoresis provides higher resolution by separating proteins based on isoelectric point (pI). This increased resolution allows researchers to distinguish between proteins that are close in molecular weight, particularly in the case of isoforms that may have differing allergenicity. The 2D gel electrophoresis technique also has low technical variability, making it a good option for research that has human health implications, such as allergen identification.4
2-D Gel Electrophoresis for Food Allergy Immunodetection
The 2D gel electrophoresis technique and subsequent immunodetection has been used to identify a variety of allergenic proteins from diverse sources. Plant pollen researchers have used this technique to identify novel IgE binding proteins from red oak pollen as well as to confirm clinically relevant allergens in a natural grass extract, including multiple isoforms of one allergenic protein.5,6 2D gel electrophoresis was used in a deep dive investigation of the major peanut allergen families, comparing different peanut strains, and has also been used to confirm no differences in allergens between natural and genetically modified soybeans.7,8
Here, we demonstrate the utility of the Auto2D® gel electrophoresis system in detecting allergens from four food sources: three plants (soybean, walnut, and buckwheat) and one animal source (salmon roe). Additionally, our system was successfully used to identify specific allergens following immunodetection using patient serum. We used the Auto2D® system to resolve single bands from SDS-PAGE into multiple distinct protein spots corresponding with major known allergens in the soybean and salmon roe samples. These results suggest that the Auto2D® gel electrophoresis system can be used for allergen identification, reducing both time and necessary technical expertise while still providing high-quality, reliable results.
Auto2D® 2-D for Food Allergy Applications Results
Soybean extract was obtained by soaking 2.5 g of commercially available soybeans in 25 mL of distilled water overnight, crushing them with a mill, and squeezing them with gauze. The protein concentration was similar to soy milk (approx. 30 µg/μL). Then, 30 μg of sample was subjected to analysis using the Auto2D® system and 0.5 μL to conventional SDS-PAGE. For 2D-E with the Auto2D® system, IEF Chip pH3-10NL was selected as the 1st-Dimension and PAGE Chip 12.5% as the 2nd-Dimension. The sample was run using the standard program (pH3-10NL M). After electrophoresis, proteins in 1D SDS-PAGE and Auto2D® gels were stained with Coomassie Brilliant Blue Stain (CBB). A separate sample was run on our Auto2D® device and was transferred to an Immobilon® P blotting membrane via semi-dry blotting. For immunodetection, the blotting membrane was blocked with 5% NFDM in PBS-T. After blocking, soybean allergic patient serum (International Bioscience, Inc.) was diluted 20x in blocking solution, added to the membrane, and incubated with gentle agitation. After washing with PBS-T, the membrane was reacted with HRP-conjugated secondary antibody diluted in blocking solution followed by standard chemiluminescence detection.
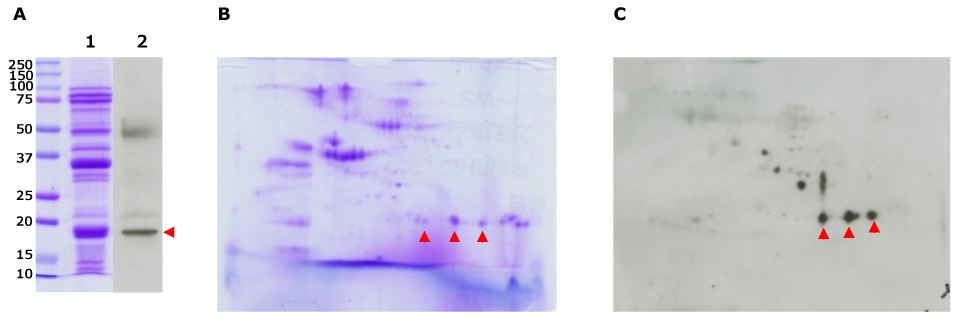
Figure 5.2D-E separation of soybean extract and immunodetection using patient serum. Soybean protein extract was separated by SDS-PAGE (A: 15 ug) and the Auto2D® system (B, C: 30 ug) using IEF Chip: pH3-10NL, PAGE Chip: 12.5% and the Tris-Glycine Reagent Kit. Whole protein in the gel was stained with CBB (A-1, B). Immunodetection was performed using soybean-allergic patient serum (A-2, C). Corresponding to the main band in SDS-PAGE, spots considered to be Gly m 6, a major soybean allergen component were detected around 18kDa (red arrows).
Each additional antigen sample was prepared under the following conditions respectively. Buckwheat flour was defatted with diethyl ether, treated with coca buffer (85 mM NaCl, 32.7 mM NaHCO3, 42.5 mM phenol), and dialyzed with PBS. To remove salts and ionic impurities that can affect the 1st-D IEF process, buckwheat extract was further processed using a protein precipitation kit (ProteoExtract® kits). Walnuts were crushed with a mortar and protein extraction was performed using the Mammalian Cell Lysis Kit (MCL1, Sigma). Salmon roe tissue was homogenized in 1M KCl-PBS. Walnut and salmon roe extracts were desalted using gel-filtration spin columns. Depending on the desalting method, protein samples were dissolved in or exchanged to rehydration solution (8M urea, 2M thiourea, 4% CHAPS, 50mM DTT, 0.02% ampholyte).
Protein quantitation for each prepared sample was performed using BCA or Bradford protein assay. Protein separation was performed using the pre-installed standard Auto2D® program (“pH3-10NL M” recipe), or by conventional SDS-PAGE. IEF Chip pH3-10NL and PAGE Chip 12.5% were selected as the 1st-D and 2nd-D gel chip to cover the widest possible separation range. After electrophoresis, separated proteins in Auto2D® gels were stained with Coomassie ReadyBlue™ stain or SYPRO® Ruby Gel stain. Separately prepared Auto2D® gels for immunodetection were removed from the chip and transferred to an Immobilon®-P blotting membrane using a wet transfer device (KS8452, Oriental instrument). PVDF membranes with transferred proteins were blocked with SuperBlock™ Blocking Buffer in PBS for 1hr. Patient sera were diluted 30-fold with PBS containing 0.1% BRIJ and allowed to react with the membranes while shaking at 4°C overnight. After washing the membrane with the above dilution buffer, alkaline phosphatase (AP) conjugated Anti-human IgE Ab (SeraCare) diluted 2,000-fold was applied as a secondary Ab and incubated at room temperature for 3hrs. After washing, the membrane was equilibrated in AP reaction buffer (100 mM Tris-HCl buffer pH 9.5 containing 100 mM NaCl and 5 mM MgCl2) and reacted with 1-component type of BCIP/NBT substrate (SeraCare) for chromogenic detection of the target proteins.
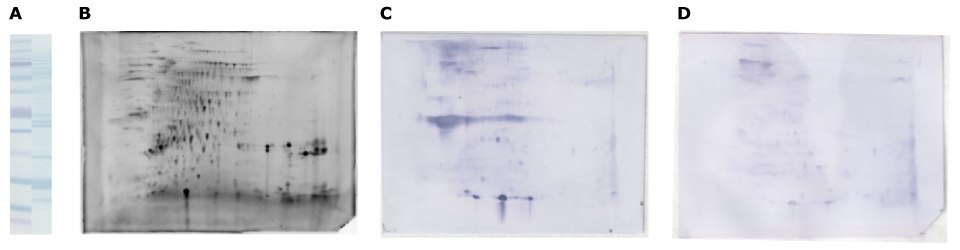
Figure 6.2D-E separation of walnut protein extract and immunodetection with allergic patient serum. Walnut protein extract was desalted in a gel-filtration spin column, equilibrated with rehydration solution, and separated by SDS-PAGE (A) and the Auto2D® system (B, C, D: 10 ug) using IEF Chip: pH3-10NL, PAGE Chip: 12.5% and Tris-Glycine Reagent Kit. Whole protein in walnut extract is visualized by amido black on membrane (A), and by SYPRO® Ruby gel stain (B), immunodetection using patient serum (C) and non-allergic serum as a negative control (D).
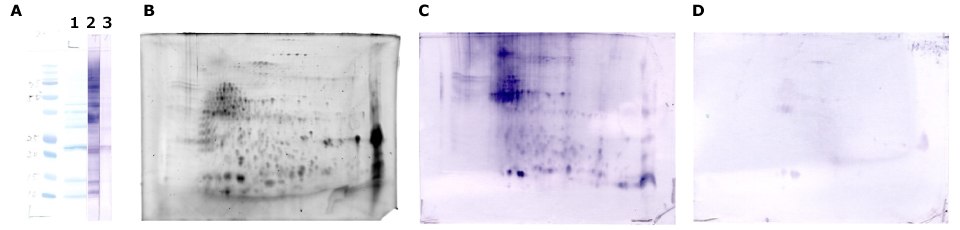
Figure 7.Immunodetection with buckwheat-allergic patient serum. Buckwheat protein extract treated with the ProteoExtract® precipitation kit was dissolved in rehydration solution and separated by SDS-PAGE (A) and the Auto2D® system (B, C, D: 10 ug) using IEF Chip: pH3-10NL, PAGE Chip: 12.5% and the Tris-Glycine Reagent Kit. Whole protein in buckwheat extract is visualized by Amido Black on membrane (A-1) or SYPRO® Ruby reagent in gel (B). Immunodetection using patient serum (A-2 and C) and non-allergic serum as a negative control (A-3 and D).
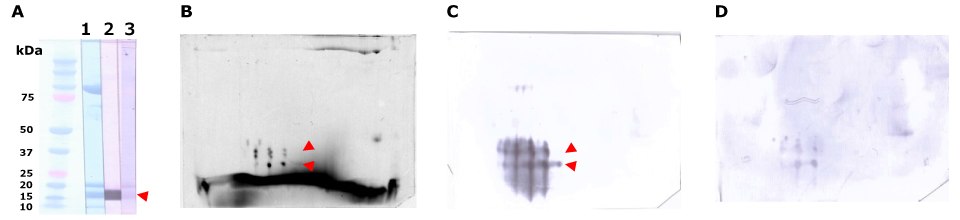
Figure 8.Separation and immunodetection of allergic components by probing with salmon roe-allergic patient serum. Salmon roe protein extract was desalted in a gel-filtration spin column, equilibrated with rehydration solution and separated by SDS-PAGE (A) and the Auto2D® system (B, C, D) using IEF Chip: pH3-10NL, PAGE Chip: 12.5%, and the Tris-Glycine Reagent Kit. Whole protein in salmon roe extract is visualized by Amido Black on membrane (A-1) or ReadyBlueTM CBB staining reagent in gel (B, 35 ug). Immunodetection using patient serum (A-2 and C, 10 ug) and non-allergic serum as a negative control (A-3 and D: 10 ug). Spots (red arrows) around 15 to 20 kDa are considered major allergens of salmon roe β’-component, which is a 35 kDa-vitellogenin-fragment consisting of two subunits.
The Western blot images were compared with total protein detection to identify the protein spots that reacted with IgE in allergic sera. To identify the allergenic component, overlapped spots are usually excised from the gel, enzymatically in-gel digested into peptide fragments, and the fragments are analyzed using mass spectral analysis. 2D-E was able to demonstrate superior resolution, showing the presence of multiple proteins in what appeared to be a single band in 1D-E. These data suggest that IgE reacting proteins can be separated and identified in fewer steps by 2D-immunodetection using allergic sera, more easily enabling the identification of causative molecules by mass spectral analysis.
Auto2D® 2-D for Food Allergy Applications Discussion
Two-dimensional gel electrophoresis is a common protein separation technique used in the detection and identification of allergenic proteins. The Auto2D® system is designed to reduce the time and technical expertise necessary to obtain high-quality results from this technique. Here, we confirm the utility of the Auto2D® system for allergy research by detecting allergens in protein samples from four food sources. Additionally, we were able to resolve a known, major allergen in each of the soybean and salmon roe samples from a single band in one-dimensional SDS-PAGE into multiple, distinct spots using the Auto2D® system. In the soybean sample, we resolved three spots from the 18kDa band that corresponds to the basic subunit of Gly m 6 (glycinin), a major, known allergenic protein.9 In the salmon roe sample, we resolved multiple spots from the 16-18kDa band that corresponds with the two subunits of β’-component (vitellogenin), also a major, known allergenic protein.10 Overall, these results suggest that the Auto2D® system can be used effectively as a fast, reliable alternative to traditional 2D gel electrophoresis for allergen immunodetection.
Acknowledgements:
We gratefully acknowledge Prof. Tatsuya Moriyama (Kindai University, Faculty of Agriculture) for providing soybean data and Prof. Yasuto Kondo (Fujita Health University Bantane Hospital) for providing walnut, buckwheat, and salmon roe data.
Simple Automated Gel Electrophoresis for Analyzing Plant Tissue-Specific Proteomes
In plant research, understanding the molecular mechanisms of physiological phenomena is a key area of study, which includes not only the analysis of individual protein functions and structures but also the analysis of protein-protein interactions and their networks. In comprehensive analyses of proteins, known as proteomics, two-dimensional electrophoresis has played an essential role in the field. While the procedures for electrophoresis itself are the same when using samples from animal tissues or microorganisms, there are significant and unique difficulties in the protein extraction or purification from plant tissues.11,12 Moreover, conventional 2D-E has challenges in learning how to operate it due to its complex and time-consuming nature, leading to data issues from lack of reproducible results.13 However, these problems are resolved by our automated system, the Auto2D® 2-D gel electrophoresis device.14
In addition, with the rapid advancements in genetics, the diversity of samples has increased, making it sometimes impossible to rely on previous studies using same/similar samples.15 Here we introduce highly versatile sample prep methods adjusted for plant sample 2-DE, using seedlings of Arabidopsis thaliana and tomato fruits, both of which are major model plants. Below, we present clear 2D separation data using the automated Auto2D® system and relevant products that are useful for protein sample prep from plant tissues.
Two Protein Extraction Methods: Direct Extraction with Auto2D® Rehydration Solution and Using A Plant Total Protein Extraction Kit
Two protein extraction methods were compared on Arabidopsis thaliana leaves. Both methods begin with grinding the frozen tissue into a powder in a cold mortar. The Plant Total Protein Extraction Kit is designed to remove polyphenolics, tannins, and other interfering substances prior to resuspending in the chaotropic reagent, which is used as the rehydration solution for the IPG strip gel.
On the other hand, extraction with the Auto2D® Rehydration Solution can be done by simple suspension of the tissue powder for high-sensitive detection such as Fluorescence or Silver, but if you need to apply a larger protein amount for Coomassie stain or MS, clean-up process using a separate kit (Protein-Concentrate Kit) will be required (recommended).
Auto2D® 2-D for Plant Proteome Applications Results
Leaf Extracts of Arabidopsis thaliana
As the result of differential fluorescence analysis, samples prepared from both extraction methods showed highly similar and acceptable 2D-E maps for downstream analysis (Figure 9A). In comparison between samples with and without Protease Inhibitor Cocktail several spots showed differences in molecular weight and pI value (Figure 9B).
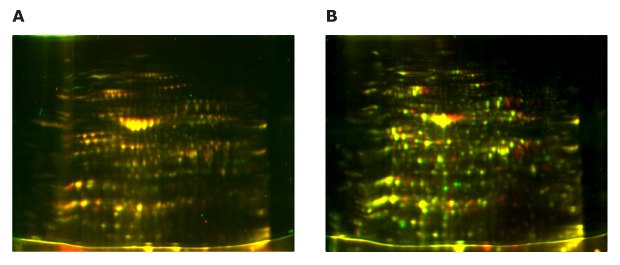
Figure 9.(A) Merged fluorescent 2D-E image of leaf extract prepared by the Plant Total Protein Extraction Kit (red) and by the combination of Auto2D® Rehydration Solution and Protein-Concentrate Kit (green). (B) Differential 2D-E image of protein extract processed in the presence (red) or absence (green) of the Protease Inhibitor Cocktail for Plant.
Organ-Specific Proteomes of Arabidopsis thaliana
Protein extracts from each organ tissue were prepared similarly to the leaf sample. Intact chloroplasts were isolated from leaves by Percoll density gradient and then dissolved in the Rehydration Solution. The protein extract from the seed was highly concentrated, so more solution can be added to suspend seed tissue powder. On the other hand, the root extract was relatively thinner, so concentration using the Protein-Concentrate Kit was effective to compare with other samples. To efficiently acquire multiple 2D-E images, all tissue extracts were labeled with fluorescent dyes here too (Figure 10).
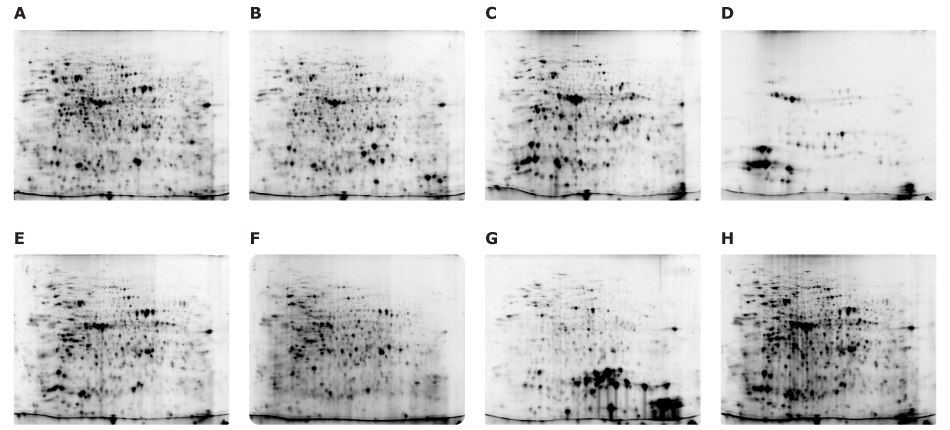
Figure 10.Fluorescence-labeled samples were analyzed in pairs using the Auto2D® device. Samples include: (A) 5 µg of Cy5 labeled flower, (B) 5 μg of Cy3 labeled seed pod, (C) 5 μg of Cy3 labeled leaf, (D) 2.5 μg of Cy5 labeled chloroplast, (E) 5 μg of Cy5 labeled stem, (F) 2.5 μg of Cy5 labeled root, (G) 5 μg of Cy3 labeled seed, and (H) 5 μg of Cy5 labeled whole-plant extract.
Suitable Plant Protein Sample Amounts For Each Detection Method
The optimum amount of protein depends on sample complexity, detection method, and its sensitivity. The Auto2D® system requires less protein amount for analysis than conventional gel electrophoresis systems. Although the loading amount should be optimized for your particular sample, detection images for each method are shown herein using the flower extract as an example (Figure 11).

Figure 11.Different amounts of flower extract samples were separated in different gels and detected with (A) 5 µg of Cy5 NHS ester, (B) 50 µg of Coomassie blue READYBLUE® stain, or (C) 3 µg of Silver Stain Kit, Sil-Best Stain One (Nacalai).
Organ-Specific Proteomes of Tomato Fruits: Sample Preparation Method for Plant Samples with High Pectin Content
Pectin is a type of polysaccharide that makes up the plant cell wall, and due to its gelling properties under acidic conditions and the presence of multiple carboxyl groups, it negatively affects 2D-E.16 However, it is difficult to remove pectin using common sample preparation methods such as Amicon® Ultra Centrifugal Filters or precipitation methods (e.g., TCA/acetone precipitation). Thus, this section will show sample preparation using pectinase with tomato fruits (Figure 12) as an example, which are a model plant abundant in pectin. Results are shown in Figure 13.

Figure 12. Diagram labeling the different organs in tomato fruits with (A) sepal, (B) pedicel, (C) inner skin, (D) seed, and (E) jelly-like tissue.
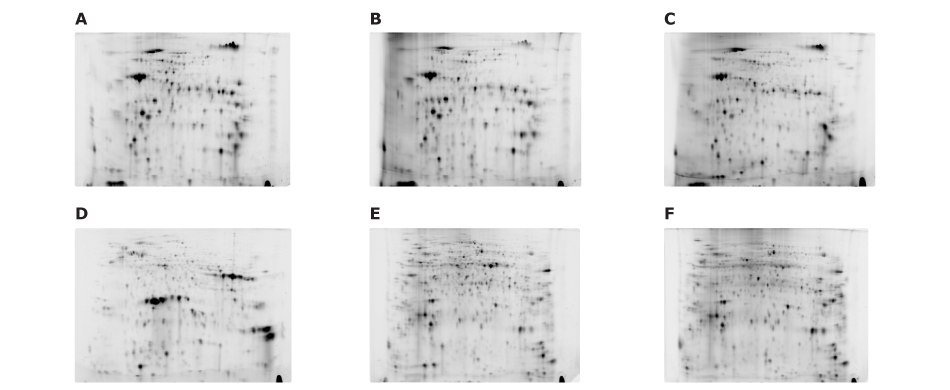

Figure 13.Fluorescence-labeled samples were analyzed in pairs using the Auto2D® device. Samples include: (A) 3 μg of Cy3 labeled juice, (B) 3 μg of Cy5 labeled inner skin, (C) 3 μg of Cy5 labeled jelly-like tissue, (D) 3 μg of Cy3 labeled seed, (E) 3 μg of Cy3 labeled sepal, and (F) 3 μg of Cy5 labeled pedicel.
Auto2d® 2-D for Plant Proteome Applications Discussion
Despite the challenges that conventional 2-DE presents, as well as the unclear guidance for plant sample prep methods, the automated Auto2D® 2-D electrophoresis device can easily, quickly, and effectively analyze plant tissues. These valuable research tools simplify 2D proteomics analysis of plant samples so that even novice electrophoresis researchers can feel confident in performing their studies.
Additionally, the sample preparation method for tomato fruits shown here may also have applications in research on food allergies and the analysis of plants as foods subject to labeling requirements. This provides those researchers with an alternative method for samples with high pectin content.
How to Obtain More Resolution When Separating Target Proteins in Protein Expression and Purification With Automated 2D Electrophoresis
In the protein purification process, SDS-PAGE is commonly used to confirm protein purity due to its simplicity. However, SDS-PAGE only allows separation based on molecular weight differences. It is not sufficient for cases that require high-purity and homogeneous samples, such as X-ray crystallography, or when evaluating heterogeneity due to post-translational modifications in antibody drugs.
On the other hand, two-dimensional electrophoresis (2D-E) allows for the separation of proteins along two axes based on differences in isoelectric point and molecular weight, making it possible to assess heterogeneity due to post-translational modifications. However, conventional 2D-E has challenges in learning how to operate it due to its complex and time-consuming nature, leading to data issues from lack of reproducible results.17
These problems are successfully resolved by our automated system, the Auto2D® 2-D electrophoresis device. This system provides quick and easy operation and reproducible results by locating difficult-to-find proteins in less than two hours. Therefore, this application note introduces the utility of 2D-E in the protein expression and purification processes using this device.
Automated 2D-E Protein Expression Results
Antibody Drugs
Monoclonal antibodies (mAbs) used in biopharmaceuticals have various heterogeneities due to differences in host cells used for antibody production, manufacturing processes, and storage conditions. These heterogeneities depend on post-translational modifications such as glycosylation, oxidation, amidation, and truncation. They can influence various biological and physicochemical properties of antibodies, including stability against proteases, half-life, and cytotoxicity.18-20 Furthermore, there are reports suggesting the possibility of involvement in the occurrence of serious side effects such as anaphylaxis and immunogenicity during the administration of antibody drugs.21,22
Currently, techniques such as mass spectrometry, SDS-PAGE, size exclusion chromatography (SEC), and capillary isoelectric focusing (cIEF) are utilized to evaluate the heterogeneity of mAb. On the other hand, 2D-E allows the comprehensive evaluation of complex mixtures of protein variants that differ in molecular weight and isoelectric point (Figure 14). Additionally, due to its separation capabilities, it can evaluate the heterogeneity of mAbs present in host cell proteins (HCPs) derived from culture supernatants (Figure 15), enabling evaluation from upstream of the manufacturing process.

Figure 14.2D-E pattern of bevacizumab under non-reduction condition. Bevacizumab was separated using the Auto2D® device under non-reduction condition. 2H2L indicates spots of complete mAb molecules composed of two heavy chains and two light chains. 2H1L, 2H, and L indicate spots of incomplete mAb molecules missing subunits.

Figure 15.2D-E pattern of mAb standard under non-reduction condition. Two mAb standard samples were separated using the Auto2D® device under non-reduction condition. (A) mAb standard, (B) mAb standard in CHO cell HCPs.
Improvement of Protein Purification Process
In X-ray crystallography, the resolution of the obtained structures depends on the quality of the protein crystals. So, it requires highly pure and homogeneous protein samples.23 As shown in Figure 16, even though the sample appears as a single band in SDS-PAGE, it was separated into three spots in the isoelectric focusing direction in 2D-E. This sample was separated into two peaks by adjusting the NaCl concentration gradient in ion exchange chromatography. The sample from peak 1 resulted in a single spot in 2D-E, allowing for the successful crystallization.
In this way, 2D-E is useful for evaluating protein homogeneity and determining directions for improving purification processes.

Figure 16.The homogeneous evaluation of sample for X-ray crystal structure analysis. (A) SDS-PAGE and (B) 2D-E pattern of partial purified protein, 2D-E patterns of (C) peak 1 and (D) peak 2 in ion-exchange column chromatography. (E) Chromatogram of ion-exchange column chromatography. Blue and red lines show UV absorption at 280 and 260 nm, respectively. Brown and green lines show the actual and programmed NaCl concentration, respectively. (F) Protein crystal obtained from peak 1. Pink and purple arrows in B, C, and D indicate spots with the same pI, respectively.
Protein Complex
In the study of protein complexes, it can be challenging to separate subunits using SDS-PAGE when their molecular weights are similar. For example, the 20S proteasome is a supramolecular complex of 750 kDa composed of 28 hetero-subunits, with each subunit concentrated in the molecular weight range of 21 to 32 kDa.24,25 This makes it difficult to separate all subunits using SDS-PAGE.
In such cases, 2D-E is useful as it can separate subunits with similar molecular weights (Figure 17). Additionally, it is capable of separating differences due to post-translational modifications, allowing for the analysis of variations in subunit phosphorylation resulting from differences in purification conditions (Figure 18).

Figure 17.SDS-PAGE and 2D-E pattern of 20S proteasome. (A) 12% SDS-PAGE, (B) 2D-E using the Auto2D® device with the proteasome subunits labeled in the pattern.
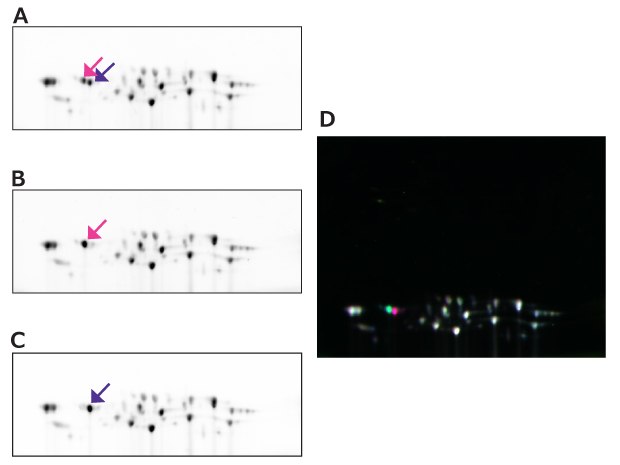

Figure 18.2D-DIGE between 20S proteasome samples purified under different conditions. (A) Cy2 labeled 20S proteasome 1, (B) Cy3 labeled 20S proteasome 2, (C) Cy5 labeled 20S proteasome 3, (D) merged image. Pink and purple arrows in A, B, and C indicate spots at the same coordinates, respectively. The spot with pink arrow and the spot with purple arrow indicate differences in the phosphorylation states of the subunits.
The Evaluation of Post-Translational Modifications In Expression Condition Screening
Post-translational modifications (PTMs) such as phosphorylation and glycosylation are closely related to protein function. When producing recombinant proteins, cell lines and expression conditions can lead to differences in these PTMs.26 Therefore, it is important to evaluate these differences and decide the expression conditions to optimize protein functionality.
The 14-3-3 protein binds to many phosphorylated proteins and is deeply involved in signal transduction. It is also known to undergo numerous PTMs itself.27 Through techniques such as 2D-DIGE and 2D-Western blotting, 2D-E allows for the detection of differences in PTMs of the target 14-3-3 protein α/β, even in crude samples like culture supernatants (Figure 19). The Auto2D® device allows for easy implementation of 2D-E, making it applicable in expression condition screening and similar applications.

Figure 19.2D-E pattern of recombinant 14-3-3 protein α/β in HEK293 cell supernatant. (A) Cy3 recombinant 14-3-3 protein α/β in HEK293 cell supernatant, (B) Cy5 control HEK293 cell supernatant, (C) merged image, (D) Cy5 recombinant 14-3-3 protein α/β in HEK293 cell supernatant on membrane, (E) Western blotting using anti-His tag antibody. The circle in A and D indicates spots derived from recombinant 14-3-3 protein α/β.
Automated 2D-E Protein Expression Discussion
It is clear that higher separation and resolution is needed to further elucidate findings in protein expression and purification processes. Techniques such as mass spectrometry, SDS-PAGE, SEC, and cIEF can provide some challenges with protein separation due to differences in protein complexes or PTMs, or needing highly purified samples for effective analysis. Here we demonstrate the utility of 2D-E in separating target proteins for expression and purification processes.
Furthermore, the data presented specifically highlights the added benefit of how the Auto2D® 2D-E device can easily, quickly, and effectively provide more resolution. With this automated device, researchers can avoid the difficulties with gel handling, extensive time, and low reproducibility that comes with traditional 2D-E. The Auto2D® device provides researchers a convenient way to separate target proteins with high reproducibility in less than 2 hours.
Residual HCP Measurement in Biopharmaceuticals
2-D gel electrophoresis is also suitable for contaminating host cell protein (HCP) analysis in therapeutic protein and antibody bioprocessing. Residual HCP impurities introduced during biologics manufacturing processes can trigger immunogenic responses in patients and reduce drug efficacy. Regulatory compliance calls for the monitoring and removal of HCPs to acceptably low levels to ensure safety. Pharmacopoeias in Japan, Europe, and the United States have published guidelines for the development and validation of HCP assays for biopharmaceutical product manufacturing.
To meet compliance, HCP assays must be methodically developed with rigorous qualification of all assay components. For antibody validation, two dimensional polyacrylamide gel electrophoresis (2D-PAGE) is required to ensure comprehensive HCP coverage. Gel electrophoresis methods including 2D-PAGE are also advised for the characterization of antigens used as calibration standards or immunogens for polyclonal antibody production. The Auto2D® 2-D Electrophoresis Device eliminates the need for advanced user training, facilitating implementation into bioprocess workflows to meet compliance with industry standards.
Auto2D® Systems for Anti-HCP Antibody Validation for ELISA and Other Immunoassays
ELISAs are the most common assay format for HCP analysis. This immunoassay method relies on polyclonal antibodies that recognize residual HCPs with broad range coverage. Japanese and European Pharmacopoeia specify the use of 2-D gel electrophoresis followed by Western blot analysis (2-D Western blotting) to evaluate HCP antibody coverage. U.S. Pharmacopoeia specifies the use of 2-D Western blotting or immunoaffinity purification followed by gel electrophoresis to evaluate antibody coverage.

Figure 20.20 μg of Cy3-labeled CHO HCP antigen from a commercial source was separated by 2-D gel electrophoresis on the Auto2D® system using a pH 3-10 IEF chip and 12.5% PAGE chip (A). Proteins were then transferred to membranes and analyzed by Western blotting using two different anti-HCP antibodies (B, C). Data indicate that anti-HCP antibody 1 has broader coverage of host cell proteins, compared to anti-HCP antibody 2.
High Reproducibility of the Auto2D® System
Consistent, reproducible processes are essential in biologics manufacturing and quality control. In addition to faster results and higher throughput, the fully automated Auto2D® gel electrophoresis system eliminates day-to-day and inter-operator variability in 2D-PAGE and 2D-DIGE methods, offering significantly better reproducibility for more reliable results.
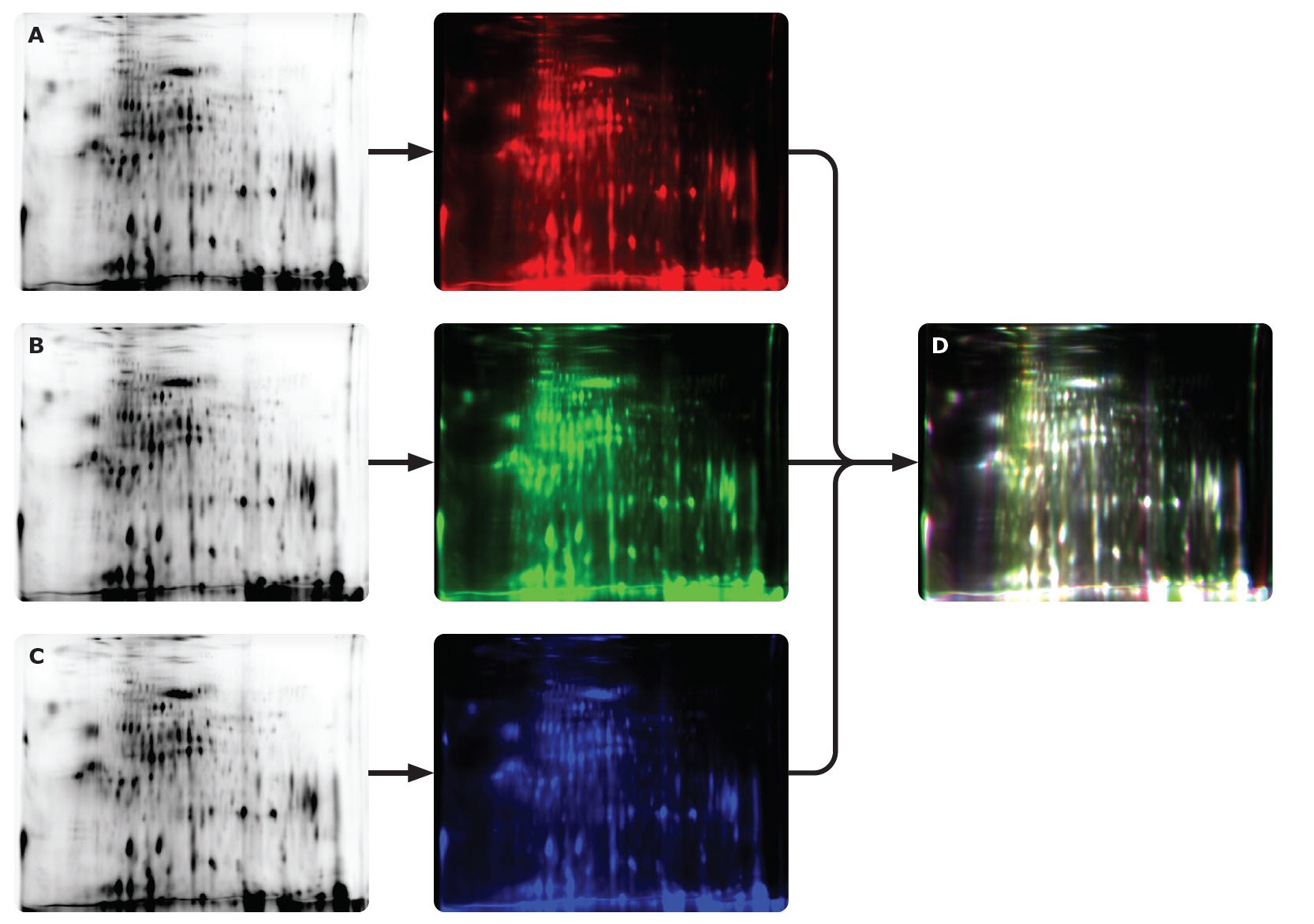
Figure 21.0.75 μg of Cy5-labeled CHO HCP antigen from a commercial source was separated by 2D gel electrophoresis and analyzed on the Auto2D® system using a IEF Chip pH 3-10NL and PAGE Chip 12.5%. The same sample was measured a total of three times (A-C). White spots (D) show overlap of results. The data suggest high reproducibility of two-dimensional gel electrophoresis using the Auto2D® system.
Auto2D® System for Antigen Profiling
Immunization with HCP antigen is required to obtain polyclonal antibody pools for ELISA and other immunoassays. HCP antigen is also used as a calibration standard for detection and quantitation. Process-specific HCP antigens are generated using non-transfected null cell cultures. These antigens must be characterized to show the pattern of HCPs, confirm the presence of a broad spectrum of proteins, and demonstrate that HCP antigens from null cultures are representative of those in production cultures. Japanese, European, and U.S. Pharmacopoeia advise the use of SDS-PAGE or 2-D gel electrophoresis for antigen characterization and profiling. U.S. Pharmacopoeia also suggests the use of 2-D gel electrophoresis as a complementary method for residual HCP monitoring.
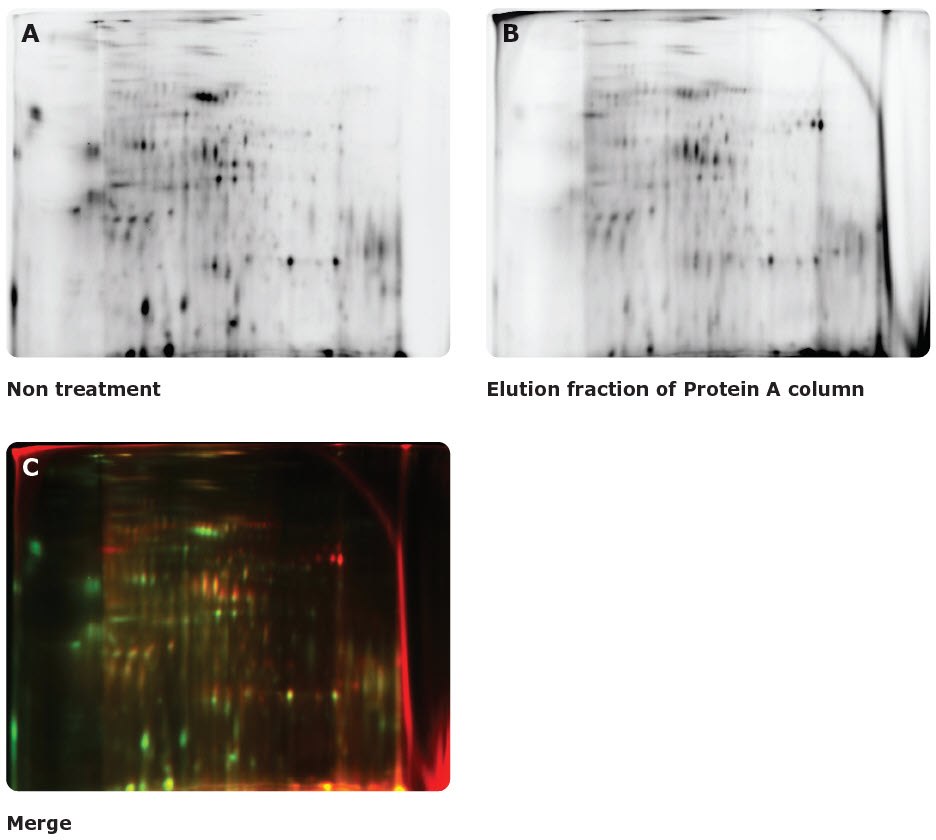
Figure 22.A mixture of Cy3-labeled unpurified CHO HCP antigen and Cy5-labeled Protein A-purified CHO HCP antigen were separated and analyzed by two-dimensional difference gel electrophoresis (2D-DIGE) using the Auto2D® system (A-B). Red spots in the merged data (C) represent host cell proteins which tend to remain even after Protein A purification. These data demonstrate the use of the Auto2D® system for profiling remaining host cell protein components post-purification.
Auto2D® System for Regulatory Compliance
The Auto2D® system helps you meet compliance with regulatory requirements for HCP analysis in biopharmaceutical manufacturing. This fully automated system eliminates day-to-day and inter-operator variability for reliable results in less than 2 hours. Easy to implement into bioprocess workflows, the Auto2D® system reduces lab downtime associated with production changes. There’s no need to outsource – protect your intellectual property using in-house resources for HCP analysis. For additional products and resources, please visit our Protein Electrophoresis and Western Blotting hub page.
TotalLab Software Enables Enhanced 2-DE with the Auto2D® 2-D Gel Electrophoresis Device
With low antibody requirements and high separation capabilities, the Auto2D® system significantly reduces the amount of time spent during sample loading, isoelectric focusing, equilibration, and SDS-PAGE. This efficiency makes the Auto2D® device unique compared to other semi-automated 2-DE systems on the market. Expanding on many of the benefits of the Auto2D® device, our collaboration with TotalLab and the coupling of their SameSpots and SpotMap software in the Auto2D® downstream workflow further enhances the previously mentioned benefits. Requiring minimal parameter setting adjustments, TotalLab software reliably demonstrates high reproducibility of spot detection when coupled with the Auto2D® device.
TotalLab Software Offers Automatic Spot Detection and Quantification
Enhanced automation and high reproducibility are achieved with the combination of the Auto2D® device and TotalLab software which improves the reproducibility and speed of 2DE and subsequent Western blot analyses. Using DIGE as the 2-DE detection method, the data below suggests that spot detection via the SameSpots software is readily achieved with samples run on gels from the fully automated Auto2D® device. Sample data is provided from gel images with spot detection results from the SameSpots software, Ver. 5.1.0.0.
The experimental materials for the data analyses include the use of extracted proteins from two types of dry yeast (Yeast A and Yeast B) using CelLytic™ Y Cell Lysis Reagent (C4482) and our Protein-Concentration Kit (2102-M). Protein was subsequently quantified with Brandford reagent (B6916) and labeled with Cy3-NHS ester or Cy5-NHS ester. Protein samples were separated on the Auto2D® device using IEF Chip pH3-10NL and 10.0% PAGE Chip while images were collected using the gel documentation system. Gels images were analyzed using the “Multiple Stains” method according to the recommended workflow analysis in the SameSpots software.
Auto2D® gel images with spot detection results from TotalLab SameSpots software, Ver. 5.1.0.0

Figure 23.The 4 sets of data for yeast extract sample data sets obtained by DIGE suggests high reproducibility from the Auto2D® device.

Figure 24.Subsequent analysis using the SameSpots software of the gels from figure 12 using the automatic alignment feature for spot matching without manual intervention (cropping/rotation/manual alignment) suggests robust and consistent spot detection.

Figure 25.Numerous spots detected by the SameSpots software with only minimal manual operations necessary.

Figure 26.Demonstration of the “Quick Tags” feature in the SameSpots software that are used to identify spots with a user-configurable fold change in expression.

Figure 27.Principal component analysis suggests strong correlation between proteins that were upregulated in the Yeast B vs the Yeast A strain.
Spot Detection Analysis of Auto2D® Gels Using TotalLab SpotMap Software Ver: 5.1.003
Alignment of spots between total protein detection on a gel and the corresponding Western Blot data is often challenging as the spot detection result is inconsistent with variable intensity for each spot. With the SpotMap software, alignment is easily achieved by the auto-align and copy alignment vectors from an aligned image functionality as highlighted with the HCP coverage assay sample data below.
HCP Coverage Assay Using the Auto2D® System and SpotMap Software
The experimental materials for the HCP coverage data analyses include the use of CHO HCP protein samples that were concentrated with our Protein-Concentration Kit (2102-M) and quantified with Bradford reagent (B6916). A fraction of the concentrated protein was labeled with Cy5-NHS ester. Approximately 2 µg of the Cy5-NHS-labeled sample and 8 µg of non-labeled sample were separated on the Auto2D® system using IEF Chip pH3-10NL, 10.0% PAGE Chip, and the gel was imaged using the gel documentation system. Proteins were subsequently transferred onto a membrane using the Immobilon®-FL (IPFL00010) by fast transfer system.
Two different anti-HCP antibodies were used for the Western blot immunodetection. HCP coverage was calculated for the gel image and Western blot image using the SpotMap software. Images were imported, and the total protein image was aligned on the membrane to the total protein image on the gel. The Western blot image was aligned to the total protein image on the gel by copying the alignment vectors. Spots were detected and assigned as either present or absent.

Figure 28.HCP coverage test with Anti-HCP antibody A. Total protein data obtained from the gel (pre-labelling, left), Western blot data with Anti-HCP antibody A (right). SpotMap version number: 5.1.003.

Figure 29.HCP coverage test with Anti-HCP antibody B. Total protein data obtained from the gel (pre-labelling, left), Western blot data with Anti-HCP antibody B (right). SpotMap version number: 5.1.003.
Compared with conventional methods, the required parameter setting adjustments were minimal and it was easy to obtain a reliable coverage percent result due to the high reproducibility of samples run on the Auto2D® device. The anti-HCP antibody coverage score was obtained in as little as 1 hour vs 12 hours using other 2D analysis software.
TotalLab Software Enhances 2-DE and Downstream Sample Analyses
The TotalLab SameSpots software automatically detects spots with high reproducibility on Auto2D® gels with no parameter settings required in advance. By combining the Auto2D® device and SameSpots software, most of the workflow steps can be automated except for image capture. The TotalLab SpotMap software can efficiently detect spots on Auto2D® gels and transfer membrane as shown in the data above, which is suitable for HCP coverage tests.
Related Products
References
To continue reading please sign in or create an account.
Don't Have An Account?